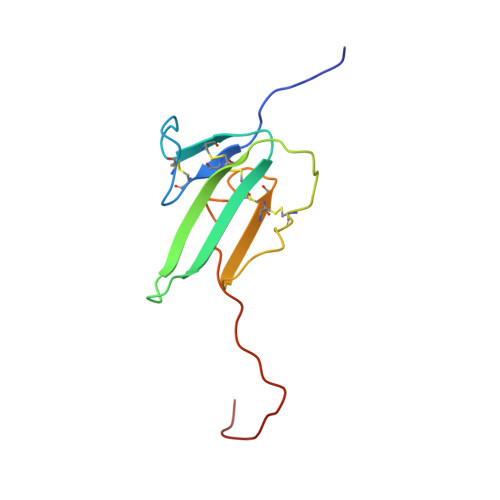
The solution structure of BMPR-IA reveals a local disorder-to-order transition upon BMP-2 binding.
Klages, J., Kotzsch, A., Coles, M., Sebald, W., Nickel, J., Muller, T., Kessler, H.(2008) Biochemistry 47: 11930-11939
- PubMed: 18937504 Search on PubMed
- DOI: https://doi.org/10.1021/bi801059j
- Primary Citation Related Structures:
2K3G - PubMed Abstract:
The structure of the extracellular domain of BMP receptor IA was determined in solution by NMR spectroscopy and compared to its structure when bound to its ligand BMP-2. While most parts of the secondary structure are highly conserved between the bound and unbound forms, large conformational rearrangements can be observed in the beta4beta5 loop of BMPR-IA, which is in contact with BMP-2 and harbors the main binding determinants for the BMPR-IA-BMP-2 interaction. In its unbound form, helix alpha1 in BMPR-IA, which is in the center of the binding epitope for BMP-2, is missing. Since BMP-2 also shows conformational changes in the type I receptor epitope upon binding to BMPR-IA, both binding partners pass through an induced fit mechanism to adapt their binding interfaces to a given interaction surface. The inherent flexibility of both partners possibly explains the promiscuous ligand-receptor interaction observed in the BMP protein superfamily.
- Center of Integrated Protein Science (CIPSM) at the Technische Universitat Munchen, Lichtenbergstrasse 4, D-85747 Garching, Germany.
Organizational Affiliation: