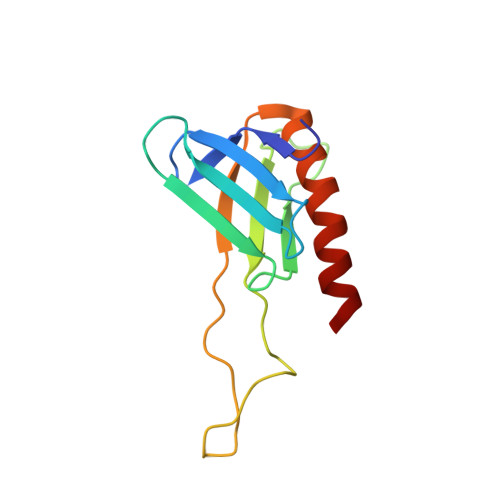
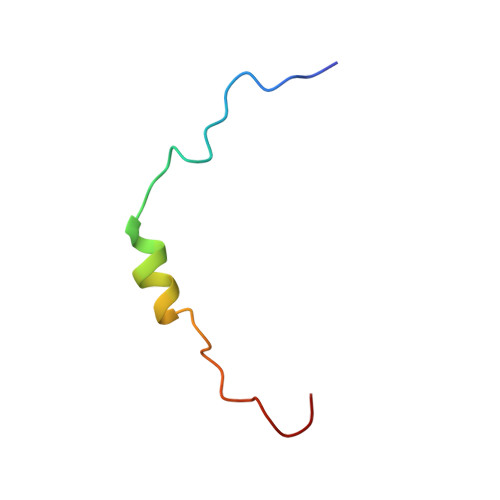
NMR Structure of the Complex between the Tfb1 Subunit of TFIIH and the Activation Domain of VP16: Structural Similarities between VP16 and p53.
Langlois, C., Mas, C., Di Lello, P., Jenkins, L.M., Legault, P., Omichinski, J.G.(2008) J Am Chem Soc 130: 10596-10604
- PubMed: 18630911
- DOI: https://doi.org/10.1021/ja800975h
- Primary Citation Related Structures:
2K2U - PubMed Abstract:
The Herpes Simplex Virion Protein 16 (VP16) activates transcription through a series of protein/protein interactions involving its highly acidic transactivation domain (TAD). The acidic TAD of VP16 (VP16TAD) has been shown to interact with several partner proteins both in vitro and in vivo, and many of these VP16 partners also bind the acidic TAD of the mammalian tumor suppressor protein p53. For example, the TADs of VP16 and p53 (p53TAD) both interact directly with the p62/Tfb1 (human/yeast) subunit of TFIIH, and this interaction correlates with their ability to activate both the initiation and elongation phase of transcription. In this manuscript, we use NMR spectroscopy, isothermal titration calorimetery (ITC) and site-directed mutagenesis studies to characterize the interaction between the VP16TAD and Tfb1. We identify a region within the carboxyl-terminal subdomain of the VP16TAD (VP16C) that has sequence similarity with p53TAD2 and binds Tfb1 with nanomolar affinity. We determine an NMR structure of a Tfb1/VP16C complex, which represents the first high-resolution structure of the VP16TAD in complex with a target protein. The structure demonstrates that like p53TAD2, VP16C forms a 9-residue alpha-helix in complex with Tfb1. Comparison of the VP16/Tfb1and p53/Tfb1 structures clearly demonstrates how the viral activator VP16C and p53TAD2 shares numerous aspects of binding to Tfb1. Despite the similarities, important differences are observed between the p53TAD2/Tfb1 and VP16C/Tfb1 complexes, and these differences demonstrate how selected activators such as p53 depend on phosphorylation events to selectively regulate transcription.
- Département de Biochimie, Université de Montréal, C.P. 6128 Succursale Centre-Ville, Montréal, QC H3C 3J7 Canada.
Organizational Affiliation: