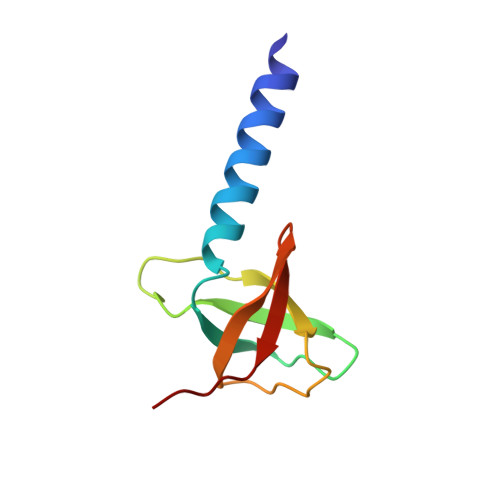
Solution structure of the soluble domain of the NfeD protein YuaF from Bacillus subtilis.
Walker, C.A., Hinderhofer, M., Witte, D.J., Boos, W., Moller, H.M.(2008) J Biomol NMR 42: 69-76
- PubMed: 18696230 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-008-9261-3
- Primary Citation Related Structures:
2K14 - PubMed Abstract:
The transmembrane protein YuaF from B. subtilis is a member of the NfeD-like clan with a potential role in maintaining membrane integrity during conditions of cellular stress. nfeD-genes are primarily found in highly conserved operon structures together with the gene of another membrane protein belonging to the SPFH superfamily, in this case YuaG. This strongly suggests a functional if not physical interaction between YuaF and YuaG. Secondary structure predictions of NfeD proteins that accompany SPFH proteins all indicate a high content of beta-sheets in the C-terminal domains indicating a conserved core structure despite very low homology at the level of primary structure. Here we report the high-resolution solution structure of YuaF's soluble C-terminal domain derived from NMR data (sYuaF, residues 97-174 of full-length YuaF). Full backbone and side chain assignments of sYuaF were obtained from triple-resonance spectra. The structure was determined from distance restraints derived from 3D NOESY spectra collected at 600 MHz and 800 MHz, together with phi, psi, and chi(1) torsion angle restraints based on the analysis of (1)H(N), (15)N, (1)H(alpha), (13)C(alpha), (13)CO, and (13)C(beta) chemical shifts, and HNHA, HNHB and HACAHB-COSY spectra. Structures were calculated using CYANA 2.0 and refined in AMBER 8. sYuaF is composed of an extended N-terminal alpha-helix and a beta-barrel formed by five beta-strands. This beta-sheet core structure is well known from the diverse class of OB-fold proteins and can also be found in the distantly related NfeD protein Ph0471 from the archaeon P. horikoshii. Despite significant differences of their amino acid sequences the structural homology of these proteins suggests a conserved function of SPFH-associated NfeD proteins.
- Department of Chemistry, University of Konstanz, Universitätsstrasse 10, 78457, Konstanz, Germany.
Organizational Affiliation: