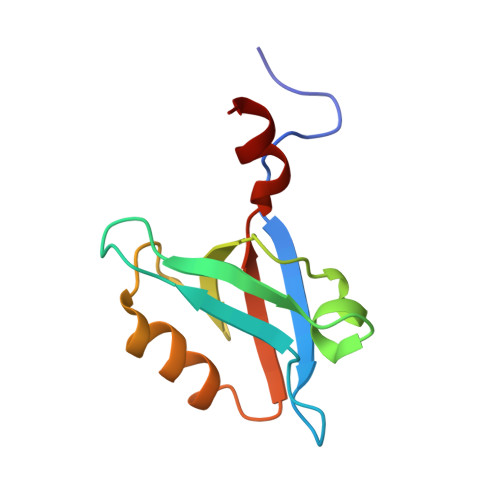
Autoinhibitory Interactions between the PDZ2 and C-terminal Domains in the Scaffolding Protein NHERF1
Cheng, H., Li, J., Fazlieva, R., Dai, Z., Bu, Z., Roder, H.(2009) Structure 17: 660-669
- PubMed: 19446522 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2009.03.009
- Primary Citation Related Structures:
2JXO - PubMed Abstract:
Na(+)/H(+) exchanger regulatory factor (NHERF1) is a signaling adaptor protein comprising two PDZ domains and a C-terminal ezrin-binding (EB) motif. To understand the role of intramolecular interactions in regulating its binding properties, we characterized the complex between the second PDZ domain PDZ2 and the C-terminal 242-358 fragment of NHERF1 using NMR and fluorescence methods. NMR chemical shift and relaxation data implicate 11 C-terminal residues in binding and, together with a thermodynamic analysis of mutant proteins, indicate that the EB region becomes helical when bound to PDZ2. Both specific contacts between PDZ2 and EB as well as nonspecific interactions involving a 100-residue flexible linker contribute to stabilizing two structurally distinct closed conformations of NHERF1. The affinity of mutant proteins for an extrinsic ligand is inversely related to the helix-forming propensity of the EB motif. The findings provide a structural framework for understanding how autoinhibitory interactions modulated the binding properties of NHERF1.
- Fox Chase Cancer Center, Philadelphia, PA 19111, USA.
Organizational Affiliation: