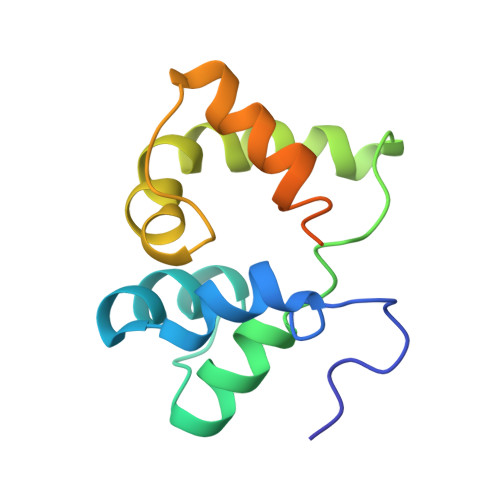
Solution NMR structure of the N-terminal domain of the human DEK protein
Devany, M., Kappes, F., Chen, K.M., Markovitz, D.M., Matsuo, H.(2008) Protein Sci 17: 205-215
- PubMed: 18227428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.073244108
- Primary Citation Related Structures:
2JX3 - PubMed Abstract:
The human DEK protein has a long-standing association with carcinogenesis since the DEK gene was originally identified in the t(6:9) chromosomal translocation in a subtype of patients with acute myelogenous leukemia (AML). Recent studies have partly unveiled DEK's cellular functions including apoptosis inhibition in primary cells as well as cancer cells, determination of 3' splice site of transcribed RNA, and suppression of transcription initiation by polymerase II. It has been previously shown that the N-terminal region of DEK, spanning residues 68-226, confers important in vitro and in vivo functions of DEK, which include double-stranded DNA (ds-DNA) binding, introduction of constrained positive supercoils into closed dsDNA, and apoptosis inhibition. In this paper, we describe the three-dimensional structure of the N-terminal domain of DEK (DEKntd) as determined using solution NMR. The C-terminal part of DEKntd, which contains a putative DNA-binding motif (SAF/SAP motif), folds into a helix-loop-helix structure. Interestingly, the N-terminal part of DEKntd shows a very similar structure to the C-terminal part, although the N-terminal and the C-terminal part differ distinctively in their amino acid sequences. As a consequence, the structure of DEKntd has a pseudo twofold plane symmetry. In addition, we tested dsDNA binding of DEKntd by monitoring changes of NMR chemical shifts upon addition of dsDNAs. We found that not only the C-terminal part containing the SAF/SAP motif but the N-terminal part is also involved in DEKntd's dsDNA binding. Our study illustrates a new structural variant and reveals novel dsDNA-binding properties for proteins containing the SAP/SAF motif.
- Department of Biochemistry, Molecular Biology and Biophysics, University of Minnesota, Minneapolis, Minnesota 55455, USA.
Organizational Affiliation: