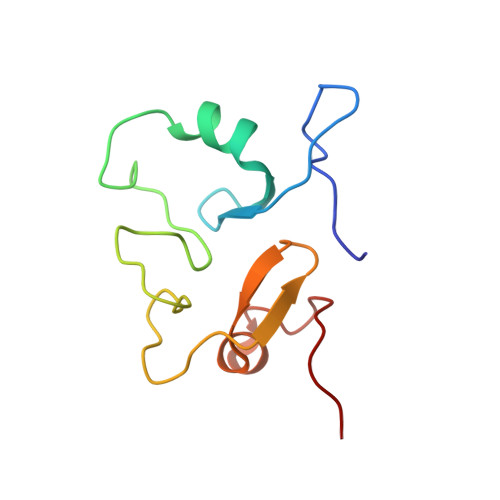
Structure of the MID1 Tandem B-Boxes Reveals an Interaction Reminiscent of Intermolecular Ring Heterodimers
Tao, H., Simmons, B.N., Singireddy, S., Jakkidi, M., Short, K.M., Cox, T.C., Massiah, M.A.(2008) Biochemistry 47: 2450-2457
- PubMed: 18220417 Search on PubMed
- DOI: https://doi.org/10.1021/bi7018496
- Primary Citation Related Structures:
2JUN - PubMed Abstract:
The tripartite motif (TRIM) protein family, defined by N-terminal RING, B-box, and coiled-coil (RBCC) domains, consists of either a single type 2 B-box domain or tandem B-box domains of type 1 and type 2 (B1B2). Here, we report the first structure of the B-box domains in their native tandem orientation. The B-boxes are from Midline-1, a putative ubiquitin E3 ligase that is required for the proteosomal degradation of the catalytic subunit of protein phosphatase 2A (PP2Ac). This function of MID1 is facilitated by the direct binding of Alpha4, a regulatory subunit of PP2Ac, to B-box1, while B-box2 appears to influence this interaction. Both B-box1 and B-box2 bind two zinc atoms in a cross-brace motif and adopt a similar betabetaalpha structure reminiscent of the RING, PHD, ZZ, and U-box domains, although they differ from each other and with RING domains in the spacing of their zinc-binding residues. The two B-box domains pack against each other with the interface formed by residues located on the structured loop consisting of the two antiparallel beta-strands. The surface area of the interface is 188 A2 (17% of the total surface). Consistent with the globular structure, the Tm of the tandem B-box domain (59 degrees C) is higher than the individual domains, supporting a stable interaction between the B-box 1 and 2 domains. Notably, the interaction is reminiscent of the interaction of recently determined RING dimers, suggesting the possibility of an evolutionarily conserved role for B-box2 domains in regulating functional RING-type folds.
- Department of Biochemistry and Molecular Biology, Oklahoma State University, Stillwater, Oklahoma 74078, USA.
Organizational Affiliation: