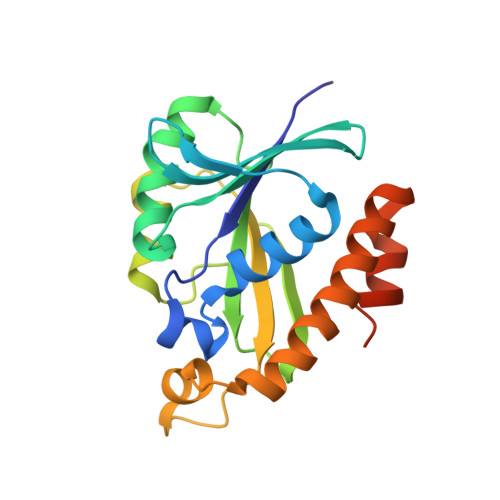
Solution structure and dynamics of peptidyl-tRNA hydrolase from Mycobacterium tuberculosis H37Rv.
Pulavarti, S.V., Jain, A., Pathak, P.P., Mahmood, A., Arora, A.(2008) J Mol Biology 378: 165-177
- PubMed: 18342886 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.02.027
- Primary Citation Related Structures:
2JRC - PubMed Abstract:
Eubacterial peptidyl-tRNA hydrolase is an essential enzyme that hydrolyzes peptidyl-tRNAs that are released into the cytoplasm because of premature termination of translation, expression of minigenes, and action of lincosamide and macrolide antibiotics. This averts the arrest of protein synthesis caused by depletion of free tRNA. Recently, we demonstrated that Mycobacterium tuberculosis peptidyl-tRNA hydrolase (MtPth) is present in the cytosol of mycobacterium and is capable of hydrolyzing peptidyl-tRNA. Here, we present the solution structure of MtPth, which is the first solution structure for this family of proteins. MtPth typically consists of seven-stranded mixed beta-sheet surrounded by six alpha-helices. The backbone dynamics for this enzyme were probed by measuring (15)N relaxation parameters and these were analyzed with model-free formalism and reduced spectral density mapping analysis. Overall, the protein molecule has tau(m) of 9.67+/-0.02 ns. The (15)N relaxation data analysis reveals that while majority of the protein backbone is rigid to motions, a short segment consisting of enzymatically critical residue H22, the loop-helix cover over the active site crevice, and the C-terminal helical hairpin exhibit motions on the milli-to microsecond timescale, all of which are linked to interaction with the substrate peptidyl-tRNA.
- Molecular and Structural Biology Division, Central Drug Research Institute, Lucknow-226 001, India.
Organizational Affiliation: