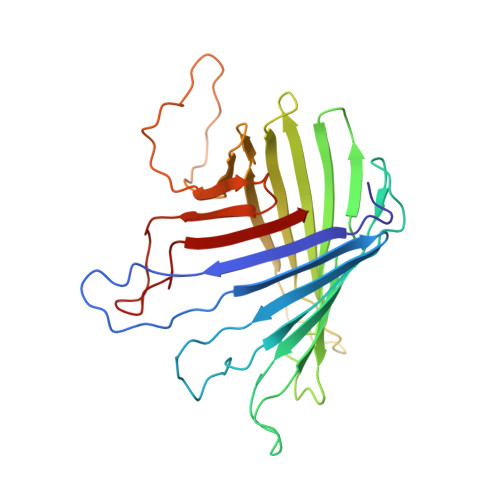
Structure of outer membrane protein G by solution NMR spectroscopy.
Liang, B., Tamm, L.K.(2007) Proc Natl Acad Sci U S A 104: 16140-16145
- PubMed: 17911261 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0705466104
- Primary Citation Related Structures:
2JQY - PubMed Abstract:
The bacterial outer membrane protein G (OmpG), a monomeric pH-gated porin, was overexpressed in Escherichia coli and refolded in beta-octyl glucoside micelles. After transfer into dodecylphosphocholine micelles, the solution structure of OmpG was determined by solution NMR spectroscopy at pH 6.3. Complete backbone assignments were obtained for 234 of 280 residues based on CA, CB, and CO connection pathways determined from a series of TROSY-based 3D experiments at 800 MHz. The global fold of the 14-stranded beta-barrel was determined based on 133 long-range NOEs observed between neighboring strands and local chemical shift and NOE information. The structure of the barrel is very similar to previous crystal structures, but the loops of the solution structure are quite flexible.
- Department of Molecular Physiology and Biological Physics, University of Virginia, 1300 Jefferson Park Avenue, PO Box 800736, Charlottesville, VA 22908, USA.
Organizational Affiliation: