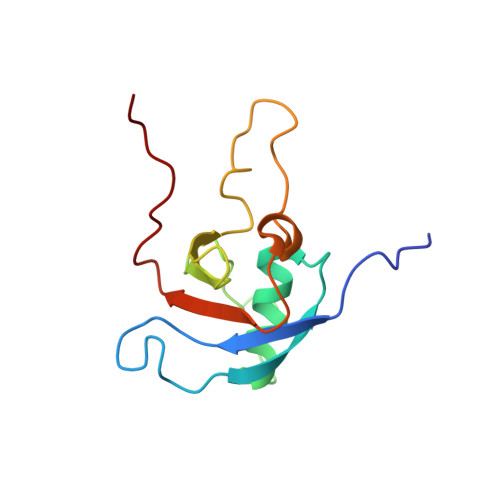
Insights into Oncogenic Mutations of Plexin-B1 Based on the Solution Structure of the Rho GTPase Binding Domain
Tong, Y., Hota, P.K., Hamaneh, M.B., Buck, M.(2008) Structure 16: 246-258
- PubMed: 18275816 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2007.12.012
- Primary Citation Related Structures:
2JPH - PubMed Abstract:
The plexin family of transmembrane receptors are important for axon guidance, angiogenesis, but also in cancer. Recently, plexin-B1 somatic missense mutations were found in both primary tumors and metastases of breast and prostate cancers, with several mutations mapping to the Rho GTPase binding domain (RBD) in the cytoplasmic region of the receptor. Here we present the NMR solution structure of this domain, confirming that the protein has both a ubiquitin-like fold and surface features. Oncogenic mutations T1795A and T1802A are located in a loop region, perturb the average structure locally, and have no effect on Rho GTPase binding affinity. Mutations L1815F and L1815P are located at the Rho GTPase binding site and are associated with a complete loss of binding for Rac1 and Rnd1. Both are found to disturb the conformation of the beta3-beta4 sheet and the orientation of surrounding side chains. Our study suggests that the oncogenic behavior of the mutants can be rationalized with reference to the structure of the RBD of plexin-B1.
- Department of Physiology and Biophysics, Case Western Reserve University School of Medicine, 10900 Euclid Avenue, Cleveland, Ohio 44106, USA.
Organizational Affiliation: