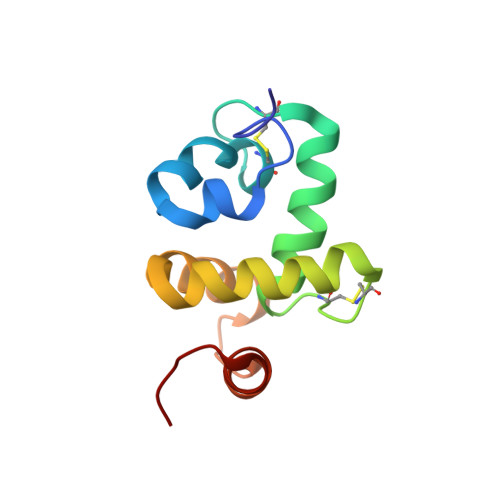
Structure of Bombyx mori chemosensory protein 1 in solution
Jansen, S., Chmelik, J., Zidek, L., Padrta, P., Novak, P., Zdrahal, Z., Picimbon, J.-F., Lofstedt, C., Sklenar, V.(2007) Arch Insect Biochem Physiol 66: 135-145
- PubMed: 17966128 Search on PubMed
- DOI: https://doi.org/10.1002/arch.20205
- Primary Citation Related Structures:
2JNT - PubMed Abstract:
Chemosensory Proteins (CSPs) represent a family of conserved proteins found in insects that may be involved in chemosensory functions. BmorCSP1 is expressed mainly in antennae and legs of the silkworm moth Bombyx mori and was cloned from antennal cDNA. Here we report the determination of the structure of Bombyx mori CSP1 (BmorCSP1) by NMR. The overall fold of BmorCSP1 is globular and comprises six alpha-helices. These helices span residues 10-14, 17-27, 35-49, 57-72, 75-85, and 92-100. The internal hydrophobic sides of the helices are formed mostly by leucine and isoleucine residues and, therefore, well suited to constitute a binding site for hydrophobic ligands.
- Department of Ecology, Lund University, Lund, Sweden.
Organizational Affiliation: