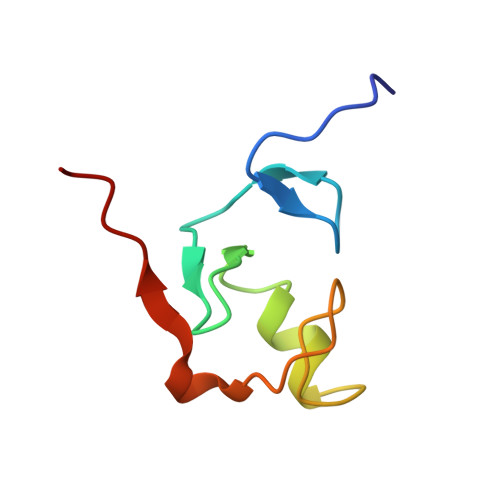
Structure, interactions, and dynamics of the RING domain from human TRAF6
Mercier, P., Lewis, M.J., Hau, D.D., Saltibus, L.F., Xiao, W., Spyracopoulos, L.(2007) Protein Sci 16: 602-614
- PubMed: 17327397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062358007
- Primary Citation Related Structures:
2JMD - PubMed Abstract:
A key step in the signaling cascade responsible for activation of the transcription factor NF-kappaB involves Lys63-linked polyubiquitination of TRAF6. Covalent attachment of ubiquitin (Ub) to TRAF6, and subsequent poly(Ub) chain synthesis, is catalyzed by the hUev1a-hUbc13 heterodimer. hUbc13 is a catalytically competent E2 enzyme, and hUev1a is an E2-like protein that binds substrate Ub. The hUev1a-hUbc13 heterodimer is targeted to TRAF6 through interactions between hUbc13 and the N-terminal RING domain from TRAF6. Nuclear magnetic resonance (NMR) spectroscopy was used to determine the solution state structure of the RING domain from human TRAF6, and the interaction between hUbc13 and TRAF6 was characterized using NMR chemical shift mapping. The main-chain dynamics of the RING domain from TRAF6 were studied using (15)N NMR relaxation. Analysis of the main-chain dynamics data indicates that residues within the alpha-helix and beta-sheet of the RING domain are as rigid as regions of canonical secondary structure in larger proteins, consistent with the biological role of RING-domain E3 proteins, which requires that the E3 contain a recognition site for recruitment of E2 ubiquitin conjugation enzymes.
- Department of Biochemistry, University of Alberta, Edmonton, Alberta T6G 2H7, Canada.
Organizational Affiliation: