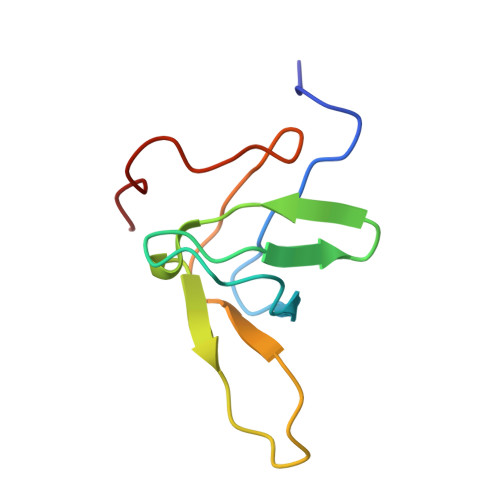
The high-resolution NMR structure of a single-chain chimeric protein mimicking a SH3-peptide complex
Candel, A.M., Conejero-Lara, F., Martinez, J.C., van Nuland, N.A.J., Bruix, M.(2007) FEBS Lett 581: 687-692
- PubMed: 17275816 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.01.032
- Primary Citation Related Structures:
2JMC - PubMed Abstract:
Here we present the high-resolution NMR structure of a chimera (SPCp41) between alpha-spectrin SH3 domain and the decapeptide p41. The tertiary structure mimics perfectly the interactions typically found in SH3-peptide complexes and is remarkably similar to that of the complex between the separate Spc-SH3 domain and ligand p41. Relaxation data confirm the tight binding between the ligand and SH3 part of the chimera. This chimera will serve as a tool for a deeper understanding of the relationship between structure and thermodynamics of binding using a combination of NMR, stability and site-directed mutagenesis studies, which can lead to an effective strategy for ligand design.
- Departamento de Química Física e Instituto de Biotecnología, Facultad de Ciencias, Universidad de Granada, Campus Fuentenueva s/n, 18071 Granada, Spain.
Organizational Affiliation: