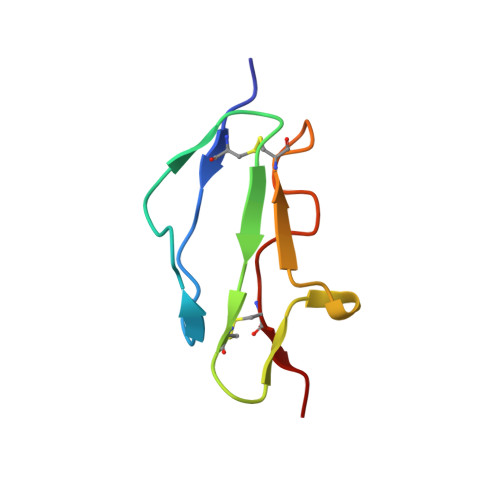
Structure shows that a glycosaminoglycan and protein recognition site in factor H is perturbed by age-related macular degeneration-linked single nucleotide polymorphism.
Herbert, A.P., Deakin, J.A., Schmidt, C.Q., Blaum, B.S., Egan, C., Ferreira, V.P., Pangburn, M.K., Lyon, M., Uhrin, D., Barlow, P.N.(2007) J Biological Chem 282: 18960-18968
- PubMed: 17360715 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M609636200
- Primary Citation Related Structures:
2JGW, 2JGX - PubMed Abstract:
A common single nucleotide polymorphism in the factor H gene predisposes to age-related macular degeneration. Factor H blocks the alternative pathway of complement on self-surfaces bearing specific polyanions, including the glycosaminoglycan chains of proteoglycans. Factor H also binds C-reactive protein, potentially contributing to noninflammatory apoptotic processes. The at risk sequence contains His (rather than Tyr) at position 402 (384 in the mature protein), in the seventh of the 20 complement control protein (CCP) modules (CCP7) of factor H. We expressed both His(402) and Tyr(402) variants of CCP7, CCP7,8, and CCP6-8. We determined structures of His(402) and Tyr(402) CCP7 and showed them to be nearly identical. The side chains of His/Tyr(402) have similar, solvent-exposed orientations far from interfaces with CCP6 and -8. Tyr(402) CCP7 bound significantly more tightly than His(402) CCP7 to a heparin affinity column as well as to defined-length sulfated heparin oligosaccharides employed in gel mobility shift assays. This observation is consistent with the position of the 402 side chain on the edge of one of two glycosaminoglycan-binding surface patches on CCP7 that we inferred on the basis of chemical shift perturbation studies with a sulfated heparin tetrasaccharide. According to surface plasmon resonance measurements, Tyr(402) CCP6-8 binds significantly more tightly than His(402) CCP6-8 to immobilized C-reactive protein. The data support a causal link between H402Y and age-related macular degeneration in which variation at position 402 modulates the response of factor H to age-related changes in the glycosaminoglycan composition and apoptotic activity of the macula.
- Edinburgh Biomolecular NMR Unit, School of Chemistry and School of Biological Sciences, University of Edinburgh, West Mains Road, Edinburgh EH9 3JJ, Scotland, United Kingdom.
Organizational Affiliation: