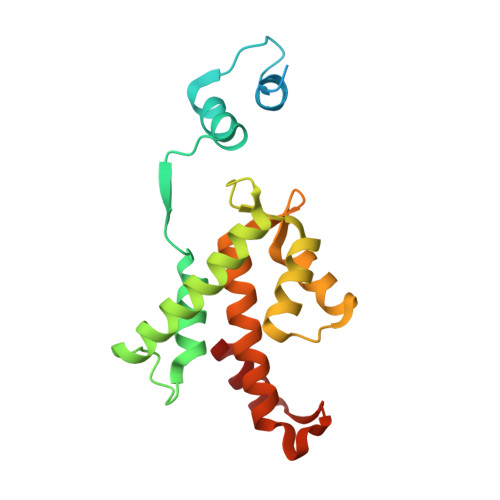
Molecular Basis for the Impaired Function of the Natural F112L Sorcin Mutant: X-Ray Crystal Structure, Calcium Affinity, and Interaction with Annexin Vii and the Ryanodine Receptor.
Franceschini, S., Ilari, A., Verzili, D., Zamparelli, C., Antaramian, A., Rueda, A., Valdivia, H.H., Chiancone, E., Colotti, G.(2008) FASEB J 22: 295
- PubMed: 17699613
- DOI: https://doi.org/10.1096/fj.07-8988com
- Primary Citation of Related Structures:
2JC2 - PubMed Abstract:
The penta-EF hand protein sorcin participates in the modulation of Ca2+-induced calcium-release in the heart through the interaction with several Ca2+ channels such as the ryanodine receptor. The modulating activity is impaired in the recently described natural F112L mutant. The F112 residue is located at the end of the D helix next to Asp113, one of the calcium ligands in the EF3 hand endowed with the highest affinity for the metal. The F112L-sorcin X-ray crystal structure at 2.5 A resolution displays marked alterations in the EF3 hand, where the hydrogen bonding network established by Phe112 is disrupted, and in the EF1 region, which is tilted in both monomers that give rise to the dimer, the stable form of the molecule. In turn, the observed tilt is indicative of an increased flexibility of the N-terminal part of the molecule. The structural alterations result in a 6-fold decrease in calcium affinity with respect to the wild-type protein and to an even larger impairment of the interaction with annexin VII and of the ability of sorcin to interact with and inhibit ryanodine receptors. These results provide a plausible structural and functional framework that helps elucidate the phenotypic alterations of mice overexpressing F112L-sorcin.
- CNR Institute of Molecular Biology and Pathology, University Sapienza, P.le A.Moro 5, 00185 Rome, Italy.
Organizational Affiliation: