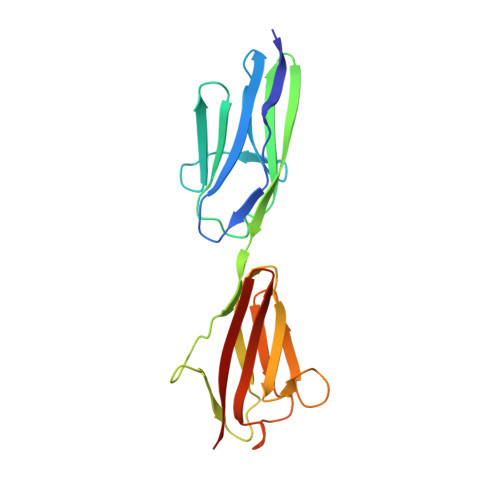
Rigid Conformation of an Immunoglobulin Domain Tandem Repeat in the A-Band of the Elastic Muscle Protein Titin
Mueller, S., Lange, S., Gautel, M., Wilmanns, M.(2007) J Mol Biology 371: 469
- PubMed: 17574571 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.05.055
- Primary Citation Related Structures:
2J8H, 2J8O - PubMed Abstract:
Most of the structure of the giant muscle protein titin is formed by small modular domains. Many of them are predicted to be arranged in repeats with short linkers that may be key determinants of the peculiar elastic properties of titin. Here, we present the molecular structure of a tandem arrangement of two immunoglobulin-like domains, A168 and A169, located within the A-band segment of titin. The two domains are connected by a 17 residue long beta-strand and form a common interface. Based on these data, we establish general principles to estimate the amount of conformational flexibility of tandem domain motifs in titin. An unusual bulge within the second domain, A169, is directly involved into binding to a sarcomeric ligand, MURF-1, thus suggesting a dual role of this tandem for both the mechanical properties of titin and for sarcomeric signaling.
- EMBL Hamburg Outstation, c/o DESY, Notkestrasse 85, D-22603 Hamburg, Germany.
Organizational Affiliation: