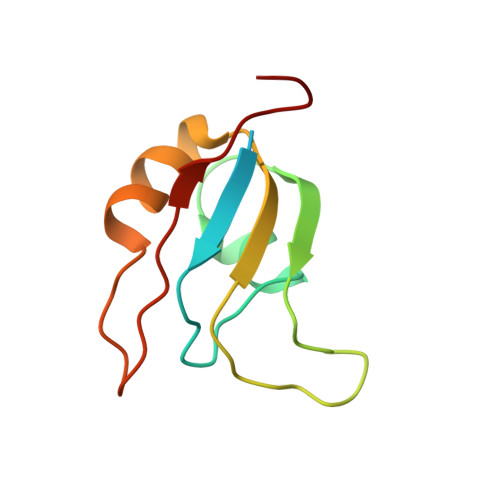
Solution Structure and RNA Interactions of the RNA Recognition Motif from Eukaryotic Translation Initiation Factor 4B.
Fleming, K., Ghuman, J., Yuan, X.M., Simpson, P., Szendroi, A., Matthews, S., Curry, S.(2003) Biochemistry 42: 8966
- PubMed: 12885229 Search on PubMed
- DOI: https://doi.org/10.1021/bi034506g
- Primary Citation Related Structures:
2J76 - PubMed Abstract:
Eukaryotic initiation factor 4B (eIF4B) is a multidomain protein with a range of activities that serves primarily to promote association of messenger RNA to the 40S ribosomal subunit during translation initiation. We report here the solution structure of the eIF4B RNA recognition motif (RRM) domain. It adopts a classical RRM fold, with a beta alpha beta beta alpha beta topology. The most striking difference with other RRM structures is in the disposition of loop 3, which connects the beta 2 and beta 3 strands and is implicated in RNA recognition. This loop folds down against the body of the RRM and exhibits restricted motion on a milli- to microsecond time scale. Although it contributes to a large basic patch on the RNA binding surface, it does not protrude out from the domain as observed in other RRM structures, possibly implying a different mode of RNA binding. On its own, the core RRM domain provides only a relative weak interaction with RNA targets and appears to require extensions at the N- and C-terminus for high-affinity binding.
- Department of Biological Sciences and Centre for Structural Biology, Imperial College London, Wolfson Laboratory, Exhibition Road, London SW7 2AY, UK.
Organizational Affiliation: