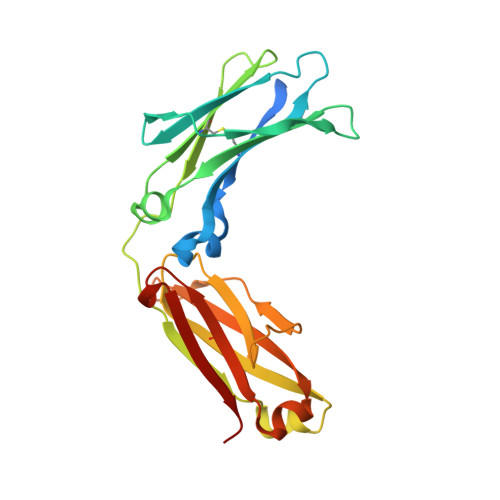
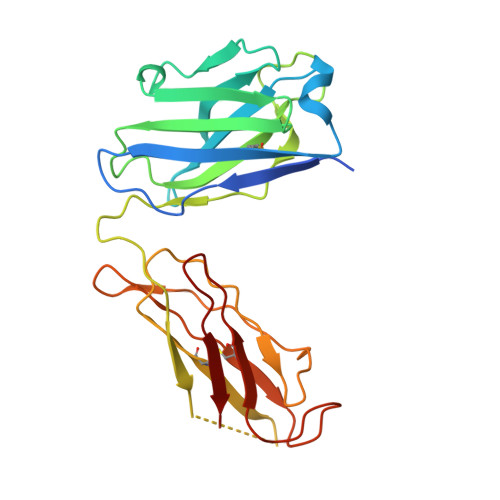
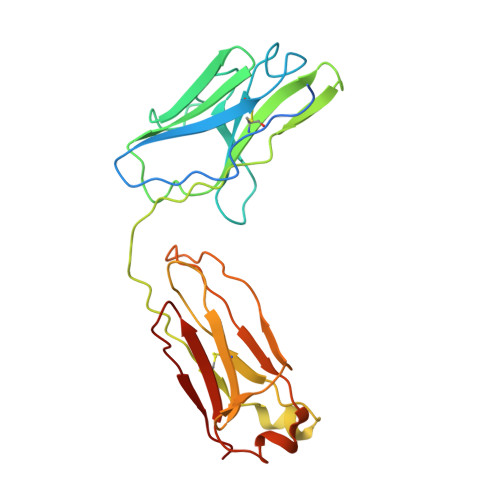
Crystal structure of a human autoimmune complex between IgM rheumatoid factor RF61 and IgG1 Fc reveals a novel epitope and evidence for affinity maturation.
Duquerroy, S., Stura, E.A., Bressanelli, S., Fabiane, S.M., Vaney, M.C., Beale, D., Hamon, M., Casali, P., Rey, F.A., Sutton, B.J., Taussig, M.J.(2007) J Mol Biology 368: 1321-1331
- PubMed: 17395205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.02.085
- Primary Citation Related Structures:
2J6E - PubMed Abstract:
Rheumatoid factors (RF) are autoantibodies that recognize epitopes in the Fc region of immunoglobulin (Ig) G and that correlate with the clinical severity of rheumatoid arthritis (RA). Here we report the X-ray crystallographic structure, at 3 A resolution, of a complex between the Fc region of human IgG1 and the Fab fragment of a monoclonal IgM RF (RF61), derived from an RA patient and with a relatively high affinity for IgG Fc. In the complex, two Fab fragments bind to each Fc at epitopes close to the C terminus, and each epitope comprises residues from both Cgamma3 domains. A central role in the unusually hydrophilic epitope is played by the side-chain of Arg355, accounting for the subclass specificity of RF61, which recognizes IgG1,-2, and -3 in preference to IgG4, in which the corresponding residue is Gln355. Compared with a previously determined complex of a lower affinity RF (RF-AN) bound to IgG4 Fc, in which only residues at the very edge of the antibody combining site were involved in binding, the epitope bound by RF61 is centered in classic fashion on the axis of the V(H):V(L) beta-barrel. The complementarity determining region-H3 loop plays a key role, forming a pocket in which Arg355 is bound by two salt-bridges. The antibody contacts also involve two somatically mutated V(H) residues, reinforcing the suggestion of a process of antigen-driven maturation and selection for IgG Fc during the generation of this RF autoantibody.
- Virologie Moléculaire et Structurale, CNRS UMR 2472-INRA UMR 1157, 91198 Gif-sur-Yvette, France.
Organizational Affiliation: