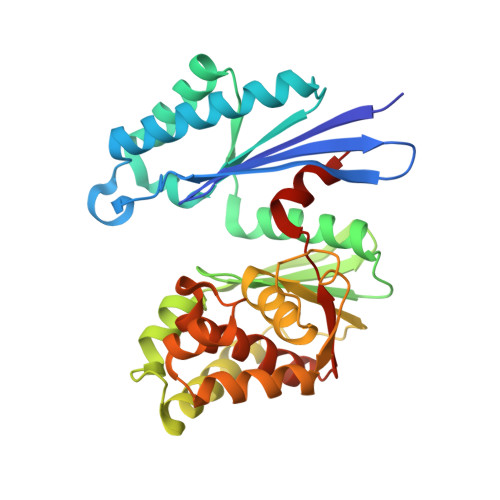
Structure of the Ppx/Gppa Phosphatase from Aquifex Aeolicus in Complex with the Alarmone Ppgpp
Kristensen, O., Ross, B., Gajhede, M.(2008) J Mol Biology 375: 1469
- PubMed: 18155044 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.11.073
- Primary Citation Related Structures:
2J4R - PubMed Abstract:
The crystal structure of the prototype exopolyphosphatase/guanosine pentaphosphate phosphohydrolase protein family member from Aquifex aeolicus in complex with the intracellular second messenger guanosine tetraphosphate was determined at 2.7-A resolution. The hydrolytic base is identified as E119. The dual specificity established for the Escherichia coli homolog is shown to be compatible with a common active site for guanosine pentaphosphate and polyphosphate hydrolysis. Distinct and different degrees of closure between the two domains of the enzyme are associated with substrate binding. The arginines R22 and R267, residing in different domains, are crucial for guanosine pentaphosphate specificity as they interact with the unique 3'-ribose phosphorylation.
- Biostructural Research, Department of Medicinal Chemistry, University of Copenhagen, Universitetsparken 2, DK-2100 Copenhagen, Denmark. ok@farma.ku.dk
Organizational Affiliation: