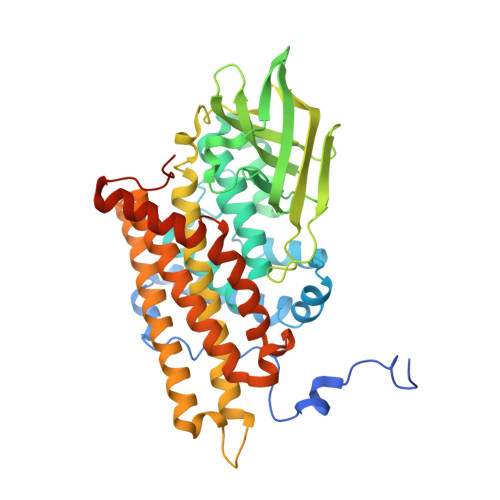
Controlling Electron Transfer in Acyl-Coa Oxidases and Dehydrogenases: A Structural View.
Mackenzie, J., Pedersen, L., Arent, S., Henriksen, A.(2006) J Biological Chem 281: 31012
- PubMed: 16887802 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M603405200
- Primary Citation Related Structures:
2IX5, 2IX6 - PubMed Abstract:
Plants produce a unique peroxisomal short chain-specific acyl-CoA oxidase (ACX4) for beta-oxidation of lipids. The short chain-specific oxidase has little resemblance to other peroxisomal acyl-CoA oxidases but has an approximately 30% sequence identity to mitochondrial acyl-CoA dehydrogenases. Two biochemical features have been linked to structural properties by comparing the structures of short chain-specific Arabidopsis thaliana ACX4 with and without a substrate analogue bound in the active site to known acyl-CoA oxidases and dehydrogenase structures: (i) a solvent-accessible acyl binding pocket is not required for oxygen reactivity, and (ii) the oligomeric state plays a role in substrate pocket architecture but is not linked to oxygen reactivity. The structures indicate that the acyl-CoA oxidases may encapsulate the electrons for transfer to molecular oxygen by blocking the dehydrogenase substrate interaction site with structural extensions. A small binding pocket observed adjoining the flavin adenine dinucleotide N5 and C4a atoms could increase the number of productive encounters between flavin adenine dinucleotide and O2.
- Biostructure Group, Carlsberg Laboratory, Gamle Carlsberg Vej 10, DK-2500 Valby, Denmark.
Organizational Affiliation: