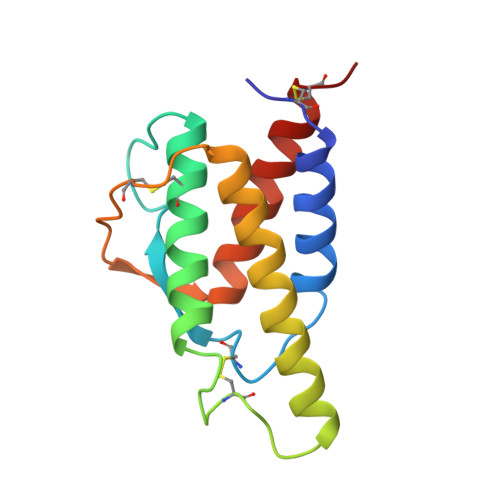
Crystal structure of recombinant human interleukin-4.
Walter, M.R., Cook, W.J., Zhao, B.G., Cameron Jr., R.P., Ealick, S.E., Walter Jr., R.L., Reichert, P., Nagabhushan, T.L., Trotta, P.P., Bugg, C.E.(1992) J Biological Chem 267: 20371-20376
- PubMed: 1400355 Search on PubMed
- DOI: https://doi.org/10.2210/pdb2int/pdb
- Primary Citation Related Structures:
2INT - PubMed Abstract:
The crystal structure of recombinant human interleukin-4 (rhuIL-4) was initially determined at 3.5-A resolution by multiple isomorphous replacement techniques and subsequently refined to a resolution of 2.35 A by simulated annealing. The final crystallographic R-factor, based on all data in the range 6.0-2.35 A (7470 reflections), is 0.232. Bond lengths and bond angles in the molecule have root mean square deviations from ideal values of 0.016 A and 2.4 degrees, respectively. The overall structure is highly compact and globular with a predominantly hydrophobic core. The main structural feature of rhuIL-4 is a four alpha-helix bundle, which composes approximately 58% of the structure. The helices are arranged in a left-handed antiparallel bundle with two overhand connections. Within these connections is a two-stranded antiparallel beta-sheet. Both the tertiary and secondary structures of rhuIL-4 are similar to those of human granulocyte-macrophage colony-stimulating factor. Critical regions for receptor binding are proposed.
- Department of Pathology, University of Alabama at Birmingham 35294.
Organizational Affiliation: