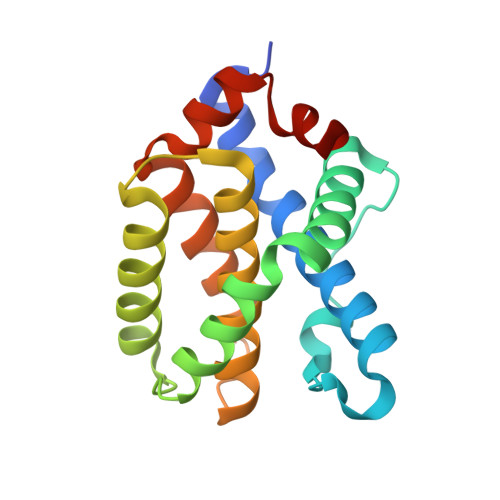
The X-ray structure of a BAK homodimer reveals an inhibitory zinc binding site.
Moldoveanu, T., Liu, Q., Tocilj, A., Watson, M., Shore, G., Gehring, K.(2006) Mol Cell 24: 677-688
- PubMed: 17157251
- DOI: https://doi.org/10.1016/j.molcel.2006.10.014
- Primary Citation Related Structures:
2IMS, 2IMT - PubMed Abstract:
BAK/BAX-mediated mitochondrial outer-membrane permeabilization (MOMP) drives cell death during development and tissue homeostasis from zebrafish to humans. In most cancers, this pathway is inhibited by BCL-2 family antiapoptotic members, which bind and block the action of proapoptotic BCL proteins. We report the 1.5 A crystal structure of calpain-proteolysed BAK, cBAK, to reveal a zinc binding site that regulates its activity via homodimerization. cBAK contains an occluded BH3 peptide binding pocket that binds a BID BH3 peptide only weakly . Nonetheless, cBAK requires activation by truncated BID to induce cytochrome c release in mitochondria isolated from bak/bax double-knockout mouse embryonic fibroblasts. The BAK-mediated MOMP is inhibited by low micromolar zinc levels. This inhibition is alleviated by mutation of the zinc-coordination site in BAK. Our results link directly the antiapoptotic effects of zinc to BAK.
- Department of Biochemistry, McGill University, 3655 Promenade Sir William Osler, Montreal, QC, Canada H3G 1Y6. Electronic address: tudor.moldoveanu@stjude.org.
Organizational Affiliation: