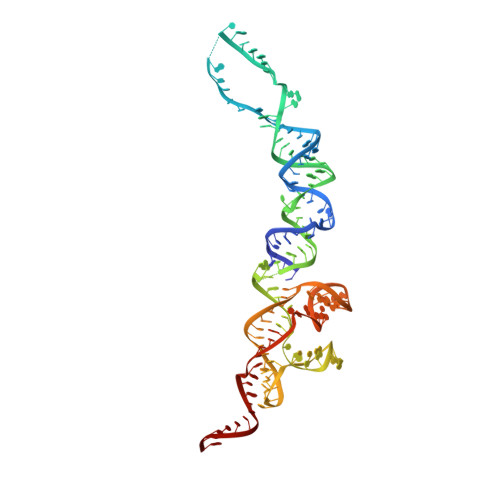
Structural basis for ribosome recruitment and manipulation by a viral IRES RNA
Pfingsten, J.S., Costantino, D.A., Kieft, J.S.(2006) Science 314: 1450-1454
- PubMed: 17124290 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1133281
- Primary Citation Related Structures:
2IL9 - PubMed Abstract:
Canonical cap-dependent translation initiation requires a large number of protein factors that act in a stepwise assembly process. In contrast, internal ribosomal entry sites (IRESs) are cis-acting RNAs that in some cases completely supplant these factors by recruiting and activating the ribosome using a single structured RNA. Here we present the crystal structures of the ribosome-binding domain from a Dicistroviridae intergenic region IRES at 3.1 angstrom resolution, providing a view of the prefolded architecture of an all-RNA translation initiation apparatus. Docking of the structure into cryo-electron microscopy reconstructions of an IRES-ribosome complex suggests a model for ribosome manipulation by a dynamic IRES RNA.
- Department of Biochemistry and Molecular Genetics, University of Colorado at Denver and Health Sciences Center, Mail Stop 8101, Post Office Box 6511, Aurora, CO 80045, USA.
Organizational Affiliation: