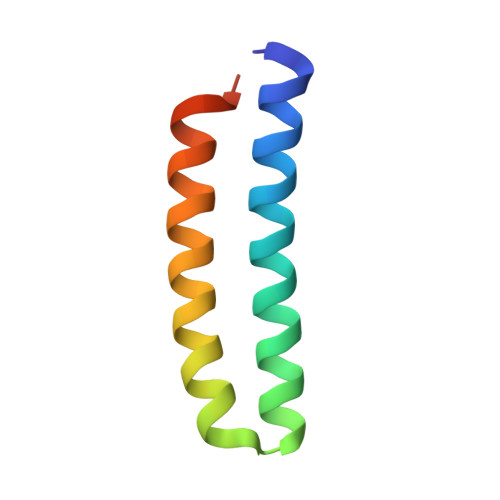
New crystal structures of ColE1 Rom and variants resulting from mutation of a surface exposed residue: Implications for RNA-recognition.
Struble, E.B., Ladner, J.E., Brabazon, D.M., Marino, J.P.(2008) Proteins 72: 761-768
- PubMed: 18260113 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21965
- Primary Citation Related Structures:
2IJH, 2IJI, 2IJJ, 2IJK - PubMed Abstract:
In ColE1, the plasmid encoded RNA one modulator (Rom) protein, which is also referred to as Rop, specifically binds and stabilizes an intermediate RNA loop-loop kissing structure formed between the plasmid encoded transcripts RNA I and RNA II and thereby acts as an auxiliary repressor of replication. Rom folds into a homodimeric, cylindrically packed four helix bundle with an exact twofold symmetry axis (Banner et al., J Mol Biol 1987;196:657-675; Eberle et al., J Biomol 1991;1:71-83). Previous studies (Castagnoli et al., EMBO J 1989;8:621-629; Predki et al., Cell 1995;80:41-50) have localized the RNA binding surface to the H1/H1' face of the helical bundle and found Phe14 to be a key determinant of the binding affinity and specificity for RNA kissing complexes. To investigate the role of Phe14 in RNA recognition, we have determined high-resolution crystal structures of two point mutants of Rom (F14Y and F14W), as well as a high-resolution structure of a crystal form of Rom in which the dimer comprises the asymmetric unit. Although the structures of F14Y and F14W share a very high degree of structural identity with that of the wild-type protein and each other, differences are observed between the three polypeptide chains found in the asymmetric unit of each crystal in the packing of the tryptophan and tyrosine side chains at position 14, as well as some of the other surface exposed side chains of key amino acids involved in RNA binding. In both the wild-type Rom and mutant structures, crystal packing forces can break the exact twofold symmetry of the dimer and influence the conformation of the side chains presented on the H1/H1' face of Rom. Since the new structures show such a high degree of structural identity, the disruption in RNA binding observed for the mutant proteins can be attributed specifically to the chemical nature of the side chain at position 14. Moreover, the fact that even subtle changes in the side chain at position 14 cannot be compensated for by the apparent flexibility of this side chain suggests a highly constrained packing of this residue in the RNA-protein complex.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, National Institute for Standards and Technology, Rockville, Maryland 20850, USA.
Organizational Affiliation: