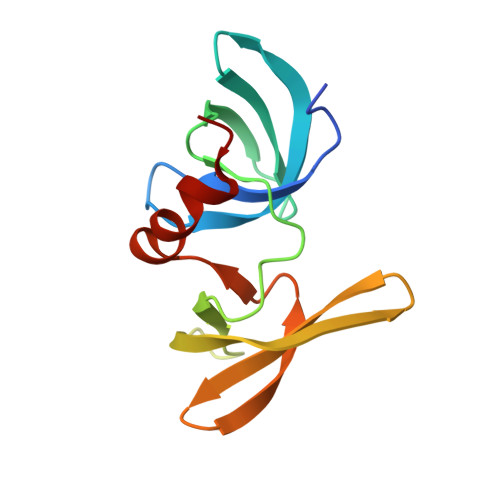
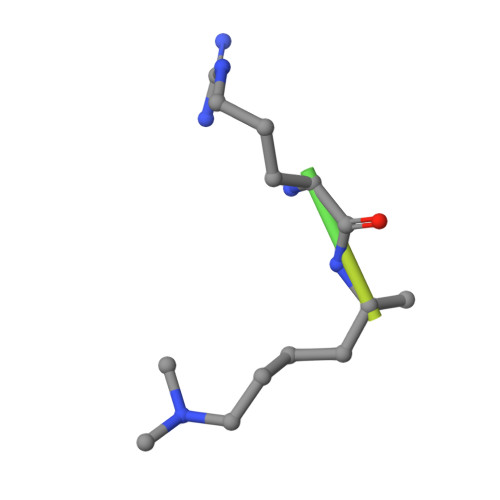
Structural Basis for the Methylation State-Specific Recognition of Histone H4-K20 by 53BP1 and Crb2 in DNA Repair.
Botuyan, M.V., Lee, J., Ward, I.M., Kim, J.E., Thompson, J.R., Chen, J., Mer, G.(2006) Cell 127: 1361-1373
- PubMed: 17190600 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2006.10.043
- Primary Citation Related Structures:
2FHD, 2G3R, 2IG0 - PubMed Abstract:
Histone lysine methylation has been linked to the recruitment of mammalian DNA repair factor 53BP1 and putative fission yeast homolog Crb2 to DNA double-strand breaks (DSBs), but how histone recognition is achieved has not been established. Here we demonstrate that this link occurs through direct binding of 53BP1 and Crb2 to histone H4. Using X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy, we show that, despite low amino acid sequence conservation, both 53BP1 and Crb2 contain tandem tudor domains that interact with histone H4 specifically dimethylated at Lys20 (H4-K20me2). The structure of 53BP1/H4-K20me2 complex uncovers a unique five-residue 53BP1 binding cage, remarkably conserved in the structure of Crb2, that best accommodates a dimethyllysine but excludes a trimethyllysine, thus explaining the methylation state-specific recognition of H4-K20. This study reveals an evolutionarily conserved molecular mechanism of targeting DNA repair proteins to DSBs by direct recognition of H4-K20me2.
- Department of Biochemistry and Molecular Biology, Mayo Clinic College of Medicine, Rochester, MN 55905, USA.
Organizational Affiliation: