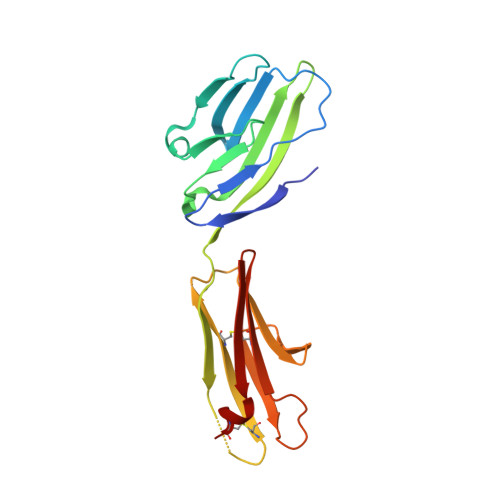NTB-A Receptor Crystal Structure: Insights into Homophilic Interactions in the Signaling Lymphocytic Activation Molecule Receptor Family.
Cao, E., Ramagopal, U.A., Fedorov, A., Fedorov, E., Yan, Q., Lary, J.W., Cole, J.L., Nathenson, S.G., Almo, S.C.(2006) Immunity 25: 559-570
- PubMed: 17045824 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2006.06.020
- Primary Citation Related Structures:
2IF7 - PubMed Abstract:
The signaling lymphocytic activation molecule (SLAM) family includes homophilic and heterophilic receptors that regulate both innate and adaptive immunity. The ectodomains of most SLAM family members are composed of an N-terminal IgV domain and a C-terminal IgC2 domain. NK-T-B-antigen (NTB-A) is a homophilic receptor that stimulates cytotoxicity in natural killer (NK) cells, regulates bactericidal activities in neutrophils, and potentiates T helper 2 (Th2) responses. The 3.0 A crystal structure of the complete NTB-A ectodomain revealed a rod-like monomer that self-associates to form a highly kinked dimer spanning an end-to-end distance of approximately 100 A. The NTB-A homophilic and CD2-CD58 heterophilic dimers show overall structural similarities but differ in detailed organization and physicochemical properties of their respective interfaces. The NTB-A structure suggests a mechanism responsible for binding specificity within the SLAM family and imposes physical constraints relevant to the colocalization of SLAM-family proteins with other signaling molecules in the immunological synapse.
- Department of Cell Biology, Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: