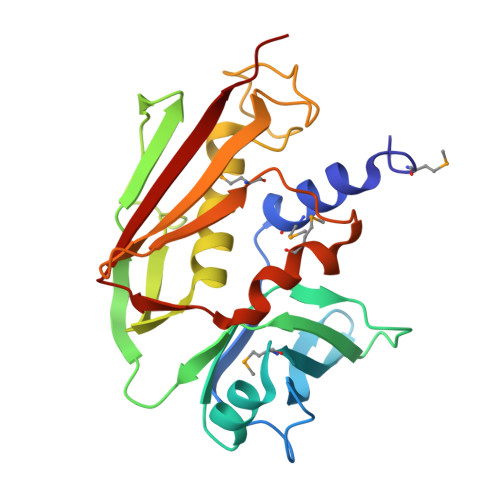
Crystal Structure of the Streptococcal Superantigen SpeI and Functional Role of a Novel Loop Domain in T Cell Activation by Group V Superantigens.
Brouillard, J.N., Gunther, S., Varma, A.K., Gryski, I., Herfst, C.A., Rahman, A.K., Leung, D.Y., Schlievert, P.M., Madrenas, J., Sundberg, E.J., McCormick, J.K.(2007) J Mol Biology 367: 925-934
- PubMed: 17303163 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.01.024
- Primary Citation Related Structures:
2ICI - PubMed Abstract:
Superantigens (SAgs) are potent microbial toxins that bind simultaneously to T cell receptors (TCRs) and class II major histocompatibility complex molecules, resulting in the activation and expansion of large T cell subsets and the onset of numerous human diseases. Within the bacterial SAg family, streptococcal pyrogenic exotoxin I (SpeI) has been classified as belonging to the group V SAg subclass, which are characterized by a unique, relatively conserved approximately 15 amino acid extension (amino acid residues 154 to 170 in SpeI; herein referred to as the alpha3-beta8 loop), absent in SAg groups I through IV. Here, we report the crystal structure of SpeI at 1.56 A resolution. Although the alpha3-beta8 loop in SpeI is several residues shorter than that of another group V SAg, staphylococcal enterotoxin serotype I, the C-terminal portions of these loops, which are located adjacent to the putative TCR binding site, are structurally similar. Mutagenesis and subsequent functional analysis of SpeI indicates that TCR beta-chains are likely engaged in a similar general orientation as other characterized SAgs. We show, however, that the alpha3-beta8 loop length, and the presence of key glycine residues, are necessary for optimal activation of T cells. Based on Vbeta-skewing analysis of human T cells activated with SpeI and structural models, we propose that the alpha3-beta8 loop is positioned to form productive intermolecular contacts with the TCR beta-chain, likely in framework region 3, and that these contacts are required for optimal TCR recognition by SpeI, and likely all other group V SAgs.
- Department of Microbiology and Immunology, The University of Western Ontario, London, ON, Canada N6A 5B8.
Organizational Affiliation: