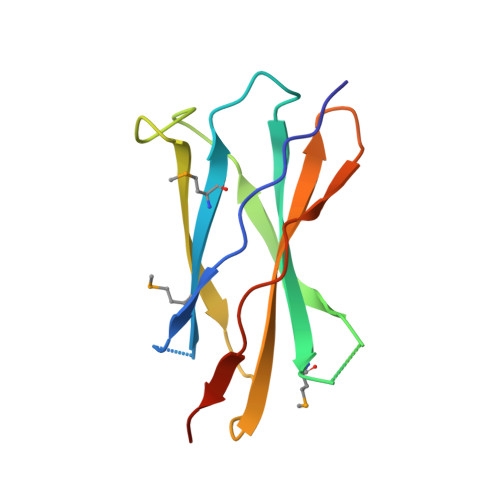
Structure of a heparin-dependent complex of Hedgehog and Ihog.
McLellan, J.S., Yao, S., Zheng, X., Geisbrecht, B.V., Ghirlando, R., Beachy, P.A., Leahy, D.J.(2006) Proc Natl Acad Sci U S A 103: 17208-17213
- PubMed: 17077139 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0606738103
- Primary Citation Related Structures:
2IBB, 2IBG, 2IC2 - PubMed Abstract:
Hedgehog (Hh) signaling molecules mediate key tissue-patterning events during animal development, and inappropriate activation of Hh signaling in adults has been associated with human cancers. Recently, a conserved family of type I integral membrane proteins required for normal response to the Hh signal was discovered. One member of this family, Ihog (interference hedgehog), functions upstream or at the level of Patched (Ptc), but how Ihog participates in Hh signaling remains unclear. Here, we show that heparin binding induces Ihog dimerization and is required to mediate high-affinity interactions between Ihog and Hh. We also present crystal structures of a Hh-binding fragment of Ihog, both alone and complexed with Hh. Heparin is not well ordered in these structures, but a basic cleft in the first FNIII domain of Ihog (IhogFn1) is shown by mutagenesis to mediate heparin binding. These results establish that Hh directly binds Ihog and provide the first demonstration of a specific role for heparin in Hh responsiveness.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: