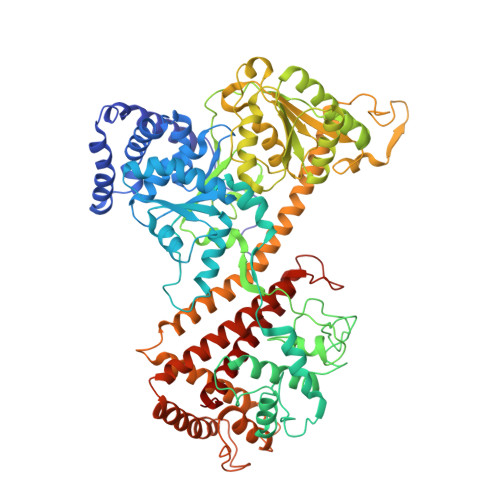
A Novel Dimer Interface and Conformational Changes Revealed by an X-ray Structure of B. subtilis SecA.
Zimmer, J., Li, W., Rapoport, T.A.(2006) J Mol Biology 364: 259-265
- PubMed: 16989859 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.08.044
- Primary Citation Related Structures:
2IBM - PubMed Abstract:
The SecA ATPase moves polypeptides post-translationally across the plasma membrane of eubacteria, but the mechanism of transport is still unclear. We describe the crystal structure of a novel dimeric form of Bacillus subtilis SecA. Dimerization of SecA occurs at the prominent groove formed by the nucleotide binding domain 2 (nbd2) and the preprotein cross-linking (ppx) domain. The dimer interface is very large, burying approximately 5400 A(2) of solvent accessible surface per monomer. Single cysteine disulfide cross-linking shows the presence of this novel SecA dimer in solution. In addition, other dimers also exist in solution, arguing that they all are in equilibrium with monomeric SecA and supporting the idea that the monomer may be the functional species. Dimerization of SecA causes an alpha-helix of one subunit to convert to a short beta-strand that participates in beta-sheet formation with strands in the other subunit. This conversion of secondary structure elements occurs close to the connection between the nbd1 and ppx domains, a potential site of interaction with translocation substrate. Comparing the different X-ray structures of B. subtilis SecA suggests that small changes in the nucleotide binding domains could be amplified via helix 1 of the helical scaffold domain (hsd) to generate larger movements of the domains involved in polypeptide binding.
- Howard Hughes Medical Institute and Department of Cell Biology, Harvard Medical School, 250 Longwood Avenue, Boston, MA 02115, USA.
Organizational Affiliation: