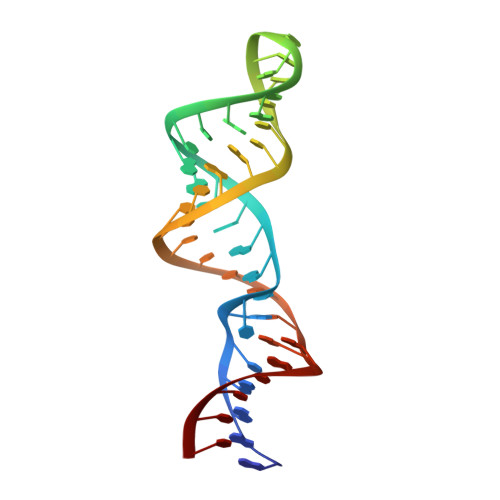
Role of metal ions in the tetraloop-receptor complex as analyzed by NMR.
Davis, J.H., Foster, T.R., Tonelli, M., Butcher, S.E.(2007) RNA 13: 76-86
- PubMed: 17119098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.268307
- Primary Citation Related Structures:
2I7E, 2I7Z - PubMed Abstract:
Metal ions are critical for the proper folding of RNA, and the GAAA tetraloop-receptor is necessary for the optimal folding and function of many RNAs. We have used NMR to investigate the role of metal ions in the structure of the tetraloop-receptor in solution. The NMR data indicate native tertiary structure is formed under a wide range of ionic conditions. The lack of conformational adaptation in response to very different ionic conditions argues against a structural role for divalent ions. Nuclear Overhauser effects to cobalt hexammine and paramagnetic relaxation enhancement induced by manganese ions were used to determine the NMR structures of the tetraloop receptor in association with metal ions, providing the first atomic-level view of these interactions in the solution state. Five manganese and two cobalt hexammine ions could be localized to the RNA surface. The locations of the associated metal ions are similar, but not identical to, those of previously determined crystal structures. The sites of association are in general agreement with nonlinear Poisson-Boltzmann calculations of the electrostatic surface, emphasizing the general importance of diffusely associated ions in RNA tertiary structure.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, Wisconsin 53706, USA.
Organizational Affiliation: