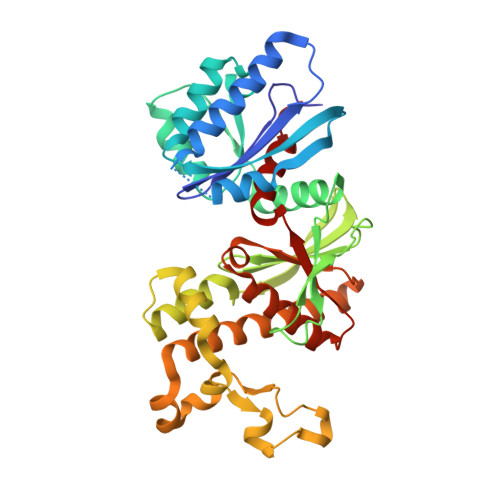
Crystal structures of human pantothenate kinases. Insights into allosteric regulation and mutations linked to a neurodegeneration disorder.
Hong, B.S., Senisterra, G., Rabeh, W.M., Vedadi, M., Leonardi, R., Zhang, Y.M., Rock, C.O., Jackowski, S., Park, H.W.(2007) J Biological Chem 282: 27984-27993
- PubMed: 17631502 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M701915200
- Primary Citation Related Structures:
2I7N, 2I7P - PubMed Abstract:
Pantothenate kinase (PanK) catalyzes the first step in CoA biosynthesis and there are three human genes that express four isoforms with highly conserved catalytic core domains. Here we report the homodimeric structures of the catalytic cores of PanK1alpha and PanK3 in complex with acetyl-CoA, a feedback inhibitor. Each monomer adopts a fold of the actin kinase superfamily and the inhibitor-bound structures explain the basis for the allosteric regulation by CoA thioesters. These structures also provide an opportunity to investigate the structural effects of the PanK2 mutations that have been implicated in neurodegeneration. Biochemical and thermodynamic analyses of the PanK3 mutant proteins corresponding to PanK2 mutations show that mutant proteins with compromised activities and/or stabilities correlate with a higher incidence of the early onset of disease.
- Structural Genomics Consortium and Department of Pharmacology, University of Toronto, Toronto, Ontario M5G 1L5, Canada.
Organizational Affiliation: