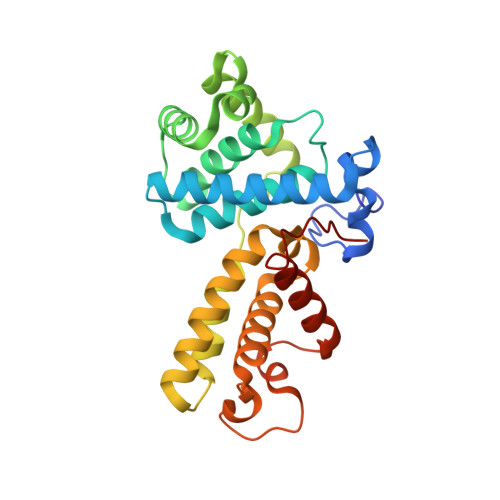
Crystal Structure of Human Cyclin K, a Positive Regulator of Cyclin-dependent Kinase 9.
Baek, K., Brown, R.S., Birrane, G., Ladias, J.A.(2007) J Mol Biology 366: 563-573
- PubMed: 17169370 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.11.057
- Primary Citation Related Structures:
2I53 - PubMed Abstract:
Cyclin K and the closely related cyclins T1, T2a, and T2b interact with cyclin-dependent kinase 9 (CDK9) forming multiple nuclear complexes, referred to collectively as positive transcription elongation factor b (P-TEFb). Through phosphorylation of the C-terminal domain of the RNA polymerase II largest subunit, distinct P-TEFb species regulate the transcriptional elongation of specific genes that play central roles in human physiology and disease development, including cardiac hypertrophy and human immunodeficiency virus-1 pathogenesis. We have determined the crystal structure of human cyclin K (residues 11-267) at 1.5 A resolution, which represents the first atomic structure of a P-TEFb subunit. The cyclin K fold comprises two typical cyclin boxes with two short helices preceding the N-terminal box. A prominent feature of cyclin K is an additional helix (H4a) in the first cyclin box that obstructs the binding pocket for the cell-cycle inhibitor p27(Kip1). Modeling of CDK9 bound to cyclin K provides insights into the structural determinants underlying the formation and regulation of this complex. A homology model of human cyclin T1 generated using the cyclin K structure as a template reveals that the two proteins have similar structures, as expected from their high level of sequence identity. Nevertheless, their CDK9-interacting surfaces display significant structural differences, which could potentially be exploited for the design of cyclin-targeted inhibitors of the CDK9-cyclin K and CDK9-cyclin T1 complexes.
- Molecular Medicine Laboratory and Macromolecular Crystallography Unit, Division of Experimental Medicine, Harvard Institutes of Medicine, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: