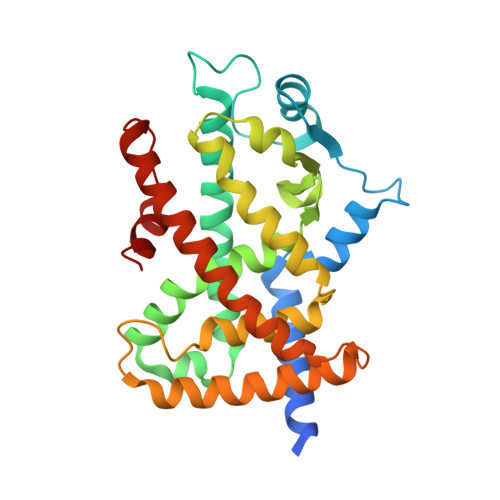
Insights into the mechanism of partial agonism: crystal structures of the peroxisome proliferator-activated receptor gamma ligand-binding domain in the complex with two enantiomeric ligands.
Pochetti, G., Godio, C., Mitro, N., Caruso, D., Galmozzi, A., Scurati, S., Loiodice, F., Fracchiolla, G., Tortorella, P., Laghezza, A., Lavecchia, A., Novellino, E., Mazza, F., Crestani, M.(2007) J Biological Chem 282: 17314-17324
- PubMed: 17403688 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M702316200
- Primary Citation Related Structures:
2I4J, 2I4P, 2I4Z - PubMed Abstract:
The peroxisome proliferator-activated receptors (PPARs) are transcriptional regulators of glucose and lipid metabolism. They are activated by natural ligands, such as fatty acids, and are also targets of synthetic antidiabetic and hypolipidemic drugs. By using cell-based reporter assays, we studied the transactivation activity of two enantiomeric ureidofibrate-like derivatives. In particular, we show that the R-enantiomer, (R)-1, is a full agonist of PPARgamma, whereas the S-enantiomer, (S)-1, is a less potent partial agonist. Most importantly, we report the x-ray crystal structures of the PPARgamma ligand binding domain complexed with the R- and the S-enantiomer, respectively. The analysis of the two crystal structures shows that the different degree of stabilization of the helix 12 induced by the ligand determines its behavior as full or partial agonist. Another crystal structure of the PPARgamma.(S)-1 complex, only differing in the soaking time of the ligand, is also presented. The comparison of the two structures of the complexes with the partial agonist reveals significant differences and is suggestive of the possible coexistence in solution of transcriptionally active and inactive forms of helix 12 in the presence of a partial agonist. Mutation analysis confirms the importance of Leu(465), Leu(469), and Ile(472) in the activation by (R)-1 and underscores the key role of Gln(286) in the PPARgamma activity.
- Istituto di Cristallografia, Consiglio Nazionale delle Ricerche, Montelibretti, 00016 Monterotondo Stazione, Roma, Italia. giorgio.pochetti@ic.cnr.it
Organizational Affiliation: