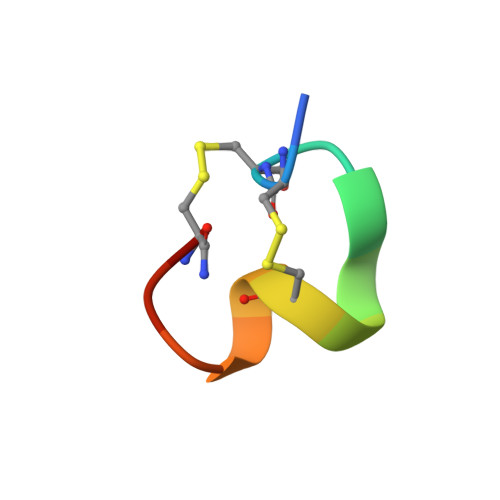
NMR structure determination of alpha-conotoxin BuIA, a novel neuronal nicotinic acetylcholine receptor antagonist with an unusual 4/4 disulfide scaffold
Chi, S.-W., Kim, D.-H., Olivera, B.M., McIntosh, J.M., Han, K.-H.(2006) Biochem Biophys Res Commun 349: 1228-1234
- PubMed: 16979596 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2006.08.164
- Primary Citation Related Structures:
2I28 - PubMed Abstract:
We have determined a high-resolution three-dimensional structure of alpha-conotoxin BuIA, a 13-residue peptide toxin isolated from Conus bullatus. Despite its unusual 4/4 disulfide bond layout alpha-conotoxin BuIA exhibits strong antagonistic activity at alpha6/alpha3beta2beta3, alpha3beta2, and alpha3beta4 nAChR subtypes like some alpha4/7 conotoxins. alpha-Conotoxin BuIA lacks the C-terminal beta-turn present within the second disulfide loop of alpha4/7 conotoxins, having only a "pseudo omega-shaped" molecular topology. Nevertheless, it contains a functionally critical two-turn helix motif, a feature ubiquitously found in alpha4/7 conotoxins. Such an aspect seems mainly responsible for similarities in the receptor recognition profile of alpha-conotoxin BuIA to alpha4/7 conotoxins. Structural comparison of alpha-conotoxin BuIA with alpha4/7 conotoxins and alpha4/3 conotoxin ImI suggests that presence of the second helical turn portion of the two-turn helix motif in alpha4/7 and alpha4/4 conotoxins may be important for binding to the alpha3 and/or alpha6 subunit of nAChR.
- Molecular Cancer Research Center, Division of Molecular Therapeutics, KRIBB, Daejeon 305-806, Republic of Korea.
Organizational Affiliation: