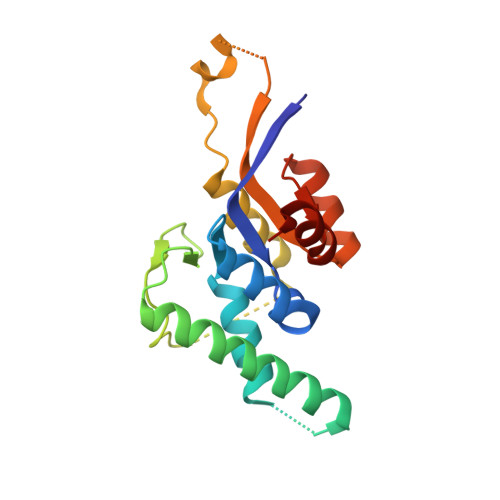
Structures of the PIN domains of SMG6 and SMG5 reveal a nuclease within the mRNA surveillance complex.
Glavan, F., Behm-Ansmant, I., Izaurralde, E., Conti, E.(2006) EMBO J 25: 5117-5125
- PubMed: 17053788 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601377
- Primary Citation Related Structures:
2HWW, 2HWX, 2HWY - PubMed Abstract:
SMG6 and SMG5 are essential factors in nonsense-mediated mRNA decay, a conserved pathway that degrades mRNAs with premature translation termination codons. Both SMG5 and SMG6 have been predicted to contain a C-terminal PIN (PilT N-terminus) domain, present in proteins with ribonuclease activity. We have determined the structures of human SMG5 and SMG6 PIN domains. Although they share a similar overall fold related to ribonucleases of the RNase H family, they have local differences at the putative active site. SMG6 has the canonical triad of acidic residues that are crucial in RNase H for nuclease activity, while SMG5 lacks key catalytic residues. The structural differences are reflected at the functional level. Only the PIN domain of SMG6 has degradation activity on single-stranded RNA in vitro. This difference in catalytic activity is conserved in Drosophila, where an SMG6 with an inactive PIN domain inhibits NMD in a dominant-negative manner. Our findings suggest that the NMD machinery has intrinsic nuclease activity that is likely to contribute to the rapid decay of mRNAs that terminate translation prematurely.
- European Molecular Biology Laboratory, Heidelberg, Germany.
Organizational Affiliation: