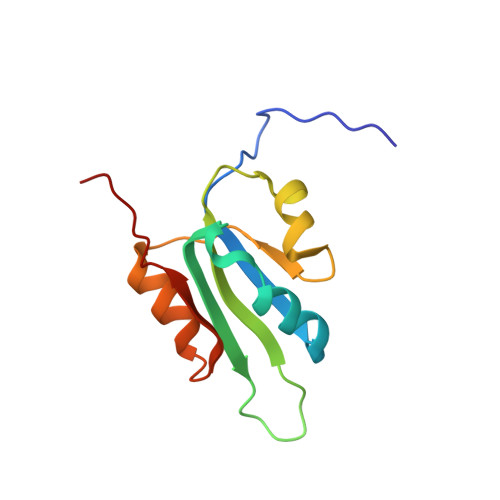
Solution structure, divalent metal and DNA binding of the endonuclease domain from the replication initiation protein from porcine circovirus 2
Vega-Rocha, S., Byeon, I.L., Gronenborn, B., Gronenborn, A.M., Campos-Olivas, R.(2007) J Mol Biology 367: 473-487
- PubMed: 17275023 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.01.002
- Primary Citation Related Structures:
2HW0 - PubMed Abstract:
Circoviruses are the smallest circular single-stranded DNA viruses able to replicate in mammalian cells. Essential to their replication is the replication initiator, or Rep protein that initiates the rolling circle replication (RCR) of the viral genome. Here we report the NMR solution three-dimensional structure of the endonuclease domain from the Rep protein of porcine circovirus type 2 (PCV2), the causative agent of postweaning multisystemic wasting syndrome in swine. The domain comprises residues 12-112 of the full-length protein and exhibits the fold described previously for the Rep protein of the representative geminivirus tomato yellow leaf curl Sardinia virus. The structure, however, differs significantly in some secondary structure elements that decorate the central five-stranded beta-sheet, including the replacement of a beta-hairpin by an alpha-helix in PCV2 Rep. The identification of the divalent metal binding site was accomplished by following the paramagnetic broadening of NMR amide signals upon Mn(2+) titration. The site comprises three conserved acidic residues on the exposed face of the central beta-sheet. For the 1:1 complex of the PCV2 Rep nuclease domain with a 22mer double-stranded DNA oligonucleotide chemical shift mapping allowed the identification of the DNA binding site on the protein and aided in constructing a model of the protein/DNA complex.
- Structural and Computational Biology Programme, Centro Nacional de Investigaciones Oncológicas, Madrid 28029, Spain.
Organizational Affiliation: