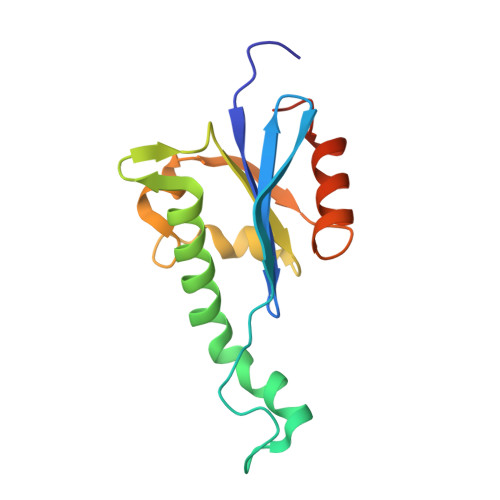
Eukaryotic class 1 translation termination factor eRF1 - the NMR structure and dynamics of the middle domain involved in triggering ribosome-dependent peptidyl-tRNA hydrolysis
Ivanova, E.V., Kolosov, P.M., Birdsall, B., Kelly, G., Pastore, A., Kisselev, L.L., Polshakov, V.I.(2007) FEBS J 274: 4223-4237
- PubMed: 17651434 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2007.05949.x
- Primary Citation Related Structures:
2HST - PubMed Abstract:
The eukaryotic class 1 polypeptide chain release factor is a three-domain protein involved in the termination of translation, the final stage of polypeptide biosynthesis. In attempts to understand the roles of the middle domain of the eukaryotic class 1 polypeptide chain release factor in the transduction of the termination signal from the small to the large ribosomal subunit and in peptidyl-tRNA hydrolysis, its high-resolution NMR structure has been obtained. The overall fold and the structure of the beta-strand core of the protein in solution are similar to those found in the crystal. However, the orientation of the functionally critical GGQ loop and neighboring alpha-helices has genuine and noticeable differences in solution and in the crystal. Backbone amide protons of most of the residues in the GGQ loop undergo fast exchange with water. However, in the AGQ mutant, where functional activity is abolished, a significant reduction in the exchange rate of the amide protons has been observed without a noticeable change in the loop conformation, providing evidence for the GGQ loop interaction with water molecule(s) that may serve as a substrate for the hydrolytic cleavage of the peptidyl-tRNA in the ribosome. The protein backbone dynamics, studied using 15N relaxation experiments, showed that the GGQ loop is the most flexible part of the middle domain. The conformational flexibility of the GGQ and 215-223 loops, which are situated at opposite ends of the longest alpha-helix, could be a determinant of the functional activity of the eukaryotic class 1 polypeptide chain release factor, with that helix acting as the trigger to transmit the signals from one loop to the other.
- Engelhardt Institute of Molecular Biology, Russian Academy of Sciences, Moscow, Russia.
Organizational Affiliation: