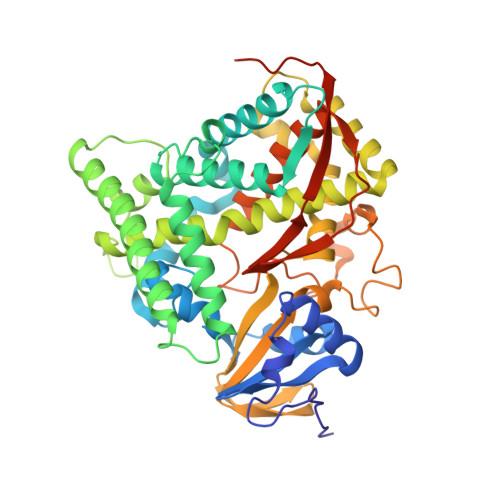
Crystal structure of hemoprotein domain of P450BM-3, a prototype for microsomal P450's.
Ravichandran, K.G., Boddupalli, S.S., Hasermann, C.A., Peterson, J.A., Deisenhofer, J.(1993) Science 261: 731-736
- PubMed: 8342039 Search on PubMed
- DOI: https://doi.org/10.1126/science.8342039
- Primary Citation Related Structures:
2HPD - PubMed Abstract:
Cytochrome P450BM-3, a bacterial fatty acid monoxygenase, resembles the eukaryotic microsomal P450's and their flavoprotein reductase in primary structure and function. The three-dimensional structure of the hemoprotein domain of P450BM-3 was determined by x-ray diffraction and refined to an R factor of 16.9 percent at 2.0 angstrom resolution. The structure consists of an alph and a beta domain. The active site heme is accessible through a long hydrophobic channel formed primarily by the beta domain and the B' and F helices of the alpha domain. The two molecules in the asymmetric unit differ in conformation around the substrate binding pocket. Substantial differences between P450BM-3 and P450cam, the only other P450 structure available, are observed around the substrate binding pocket and the regions important for redox partner binding. A general mechanism for proton transfer in P450's is also proposed.
- Howard Hughes Medical Institute, University of Texas Southwestern Medical Center, Dallas 75235-9050.
Organizational Affiliation: