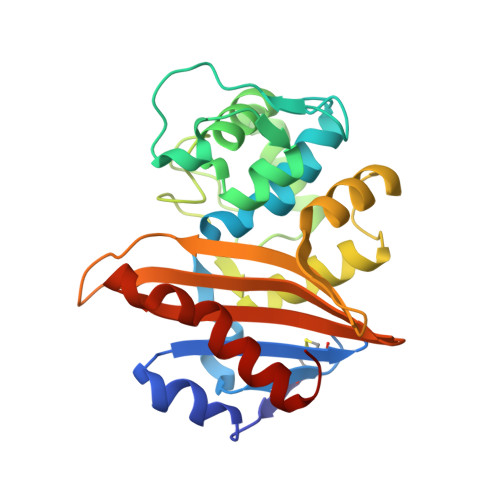
Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Baurin, S., Vercheval, L., Bouillenne, F., Falzone, C., Brans, A., Jacquamet, L., Ferrer, J.L., Sauvage, E., Dehareng, D., Frere, J.M., Charlier, P., Galleni, M., Kerff, F.(2009) Biochemistry 48: 11252-11263
- PubMed: 19860471 Search on PubMed
- DOI: https://doi.org/10.1021/bi901548c
- Primary Citation Related Structures:
2HP5, 2HP6, 2HP9, 2HPB, 2RL3, 2WGI - PubMed Abstract:
The catalytic efficiency of the class D beta-lactamase OXA-10 depends critically on an unusual carboxylated lysine as the general base residue for both the enzyme acylation and deacylation steps of catalysis. Evidence is presented that the interaction between the indole group of Trp154 and the carboxylated lysine is essential for the stability of the posttranslationally modified Lys70. Substitution of Trp154 by Gly, Ala, or Phe yielded noncarboxylated enzymes which displayed poor catalytic efficiencies and reduced stability when compared to the wild-type OXA-10. The W154H mutant was partially carboxylated. In addition, the maximum values of k(cat) and k(cat)/K(M) were shifted toward pH 7, indicating that the carboxylation state of Lys70 is dependent on the protonation level of the histidine. A comparison of the three-dimensional structures of the different proteins also indicated that the Trp154 mutations did not modify the overall structures of OXA-10 but induced an increased flexibility of the Omega-loop in the active site. Finally, the deacylation-impaired W154A mutant was used to determine the structure of the acyl-enzyme complex with benzylpenicillin. These results indicate a role of the Lys70 carboxylation during the deacylation step and emphasize the importance of Trp154 for the ideal positioning of active site residues leading to an optimum activity.
- Laboratory of Biological Macromolecules, Center for Protein Engineering, University of Liège, Institut de Chimie B6a, Belgium.
Organizational Affiliation: