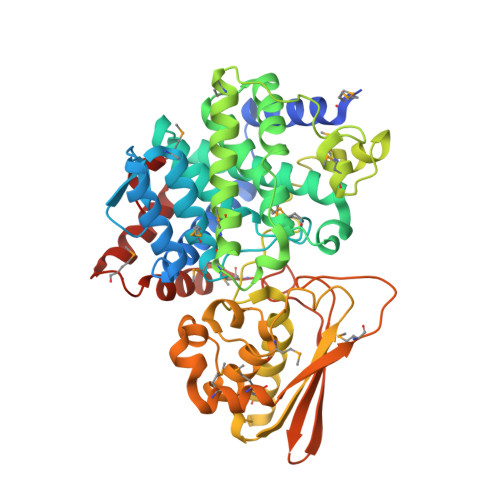
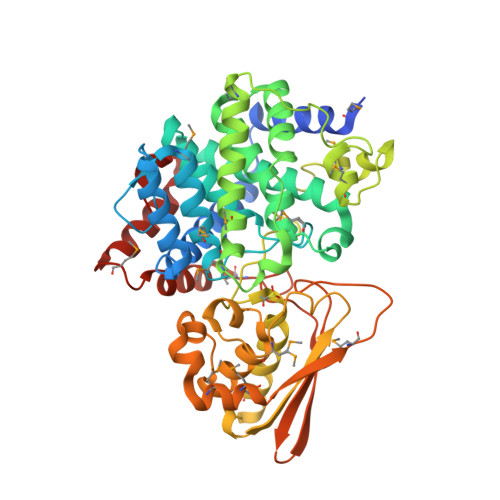
Three-dimensional Structure of Iminodisuccinate Epimerase Defines the Fold of the MmgE/PrpD Protein Family.
Lohkamp, B., Bauerle, B., Rieger, P.G., Schneider, G.(2006) J Mol Biology 362: 555-566
- PubMed: 16934291 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.07.051
- Primary Citation Related Structures:
2HP0, 2HP3 - PubMed Abstract:
Iminodisuccinate (IDS) epimerase catalyzes the epimerisation of R,R-, S,S- and R,S- iminodisuccinate, one step in the biodegradation of the chelating agent iminodisuccinate by Agrobacterium tumefaciens BY6. The enzyme is a member of the MmgE/PrpD protein family, a diverse and little characterized class of proteins of prokaryotic and eukaryotic origin. IDS epimerase does not show significant overall amino acid sequence similarity to any other protein of known three-dimensional structure. The crystal structure of this novel epimerase has been determined by multi-wavelength diffraction to 1.5 A resolution using selenomethionine-substituted enzyme. In the crystal, the enzyme forms a homo-dimer, and the subunit consists of two domains. The larger domain, not consecutive in sequence and comprising residues Met1-Lys266 and Leu400-Pro446, forms a novel all alpha-helical fold with a central six-helical bundle. The second, smaller domain folds into an alpha+beta domain, related in topology to chorismate mutase by a circular permutation. IDS epimerase is thus not related in three-dimensional structure to other known epimerases. The fold of the IDS epimerase is representative for the whole MmgE/PrpD family. The putative active site is located at the interface between the two domains of the subunit, and is characterized by a positively charged surface, consistent with the binding of a highly negatively charged substrate such as iminodisuccinate. Docking experiments suggest a two-base mechanism for the epimerisation reaction.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, 171 77 Stockholm, Sweden.
Organizational Affiliation: