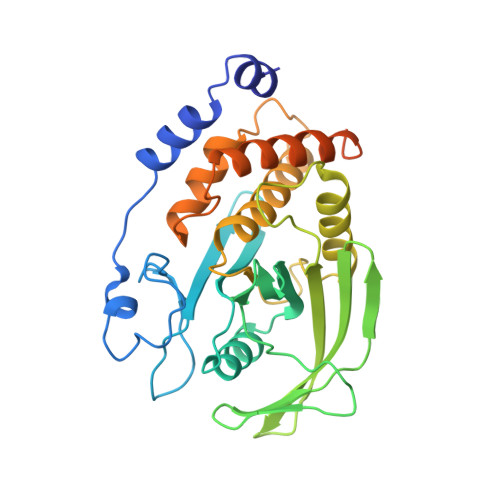
Crystal structure of human protein tyrosine phosphatase 1B.
Barford, D., Flint, A.J., Tonks, N.K.(1994) Science 263: 1397-1404
- PubMed: 8128219 Search on PubMed
- Primary Citation Related Structures:
2HNP, 2HNQ - PubMed Abstract:
Protein tyrosine phosphatases (PTPs) constitute a family of receptor-like and cytoplasmic signal transducing enzymes that catalyze the dephosphorylation of phosphotyrosine residues and are characterized by homologous catalytic domains. The crystal structure of a representative member of this family, the 37-kilodalton form (residues 1 to 321) of PTP1B, has been determined at 2.8 A resolution. The enzyme consists of a single domain with the catalytic site located at the base of a shallow cleft. The phosphate recognition site is created from a loop that is located at the amino-terminus of an alpha helix. This site is formed from an 11-residue sequence motif that is diagnostic of PTPs and the dual specificity phosphatases, and that contains the catalytically essential cysteine and arginine residues. The position of the invariant cysteine residue within the phosphate binding site is consistent with its role as a nucleophile in the catalytic reaction. The structure of PTP1B should serve as a model for other members of the PTP family and as a framework for understanding the mechanism of tyrosine dephosphorylation.
- W.M. Keck Structural Biology Laboratory, Cold Spring Harbor Laboratory, NY 11724.
Organizational Affiliation: