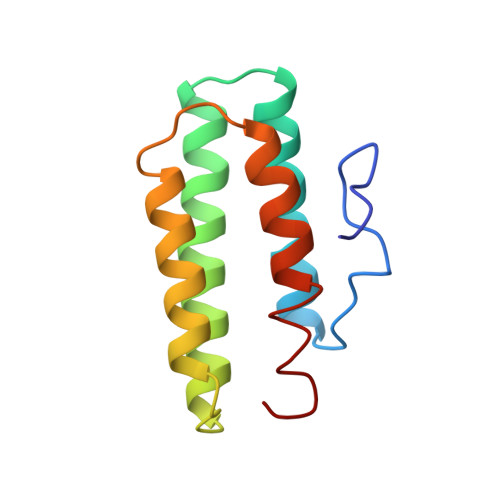
Structures of met and azidomet hemerythrin at 1.66 A resolution.
Holmes, M.A., Stenkamp, R.E.(1991) J Mol Biology 220: 723-737
- PubMed: 1870128 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90113-k
- Primary Citation Related Structures:
2HMQ, 2HMZ - PubMed Abstract:
The crystallographic refinement of met and azidomet hemerythrin has been carried out at 1.66 A resolution in an attempt to characterize precisely the binuclear iron center in this protein. Restrained least-squares refinement has produced molecular models giving R-values of 18.9% for met (65,683 reflections from 10 A to 1.66 A) and 17.6% for azidomet hemerythrin (68,747 reflections from 10.0 A to 1.66 A). The protein structure in each derivative is very similar to that of myohemerythrin. The mu-oxo bridged iron center differs between the two forms. The complex in met hemerythrin is asymmetric with the bridging oxygen closer to one of the iron atoms while the complex in azidomet hemerythrin is symmetric. After investigations of the effects of correlation in the refinement, we believe this difference between the two complexes is associated with chemical differences and is not a refinement artefact.
- Department of Biological Structure, University of Washington, Seattle 98195.
Organizational Affiliation: