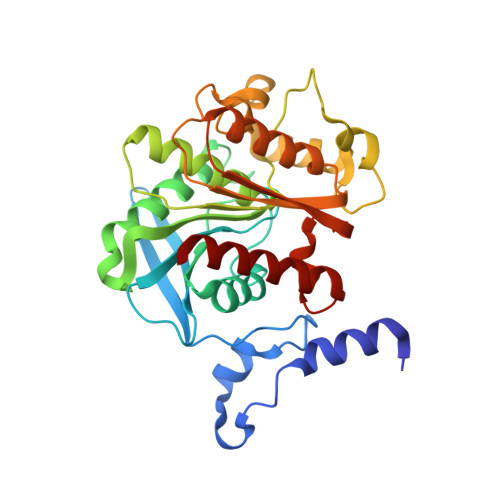
Functional and structural features of the oxyanion hole in a thermophilic esterase from Alicyclobacillus acidocaldarius.
Mandrich, L., Menchise, V., Alterio, V., De Simone, G., Pedone, C., Rossi, M., Manco, G.(2007) Proteins 71: 1721-1731
- PubMed: 18076040 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21877
- Primary Citation Related Structures:
2HM7 - PubMed Abstract:
Recent mutagenic and molecular modelling studies suggested a role for glycine 84 in the putative oxyanion loop of the carboxylesterase EST2 from Alicyclobacillus acidocaldarius. A 114 times decrease of the esterase catalytic activity of the G84S mutant was observed, without changes in the thermal stability. The recently solved three-dimensional (3D) structure of EST2 in complex with a HEPES molecule permitted to demonstrate that G84 (together with G83 and A156) is involved in the stabilization of the oxyanion through a hydrogen bond from its main chain NH group. The structural data in this case did not allowed us to rationalize the effect of the mutation, since this hydrogen bond was predicted to be unaltered in the mutant. Since the mutation could shed light on the role of the oxyanion loop in the HSL family, experiments to elucidate at the mechanistic level the reasons of the observed drop in k (cat) were devised. In this work, the kinetic and structural features of the G84S mutant were investigated in more detail. The optimal temperature and pH for the activity of the mutated enzyme were found significantly changed (T = 65 degrees C and pH = 5.75). The catalytic constants K (M) and V(max) were found considerably altered in the mutant, with ninefold increased K (M) and 14-fold decreased V(max), at pH 5.75. At pH 7.1, the decrease in k (cat) was much more dramatic. The measurement of kinetic constants for some steps of the reaction mechanism and the resolution of the mutant 3D structure provided evidences that the observed effects were partly due to the steric hindrance of the S84-OH group towards the ester substrate and partly to its interference with the nucleophilic attack of a water molecule on the second tetrahedral intermediate.
- Istituto di Biochimica delle Proteine, CNR, Via P. Castellino 111, 80131 Naples, Italy.
Organizational Affiliation: