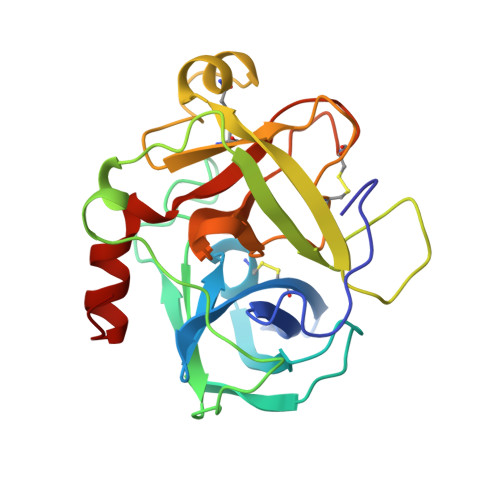
1.7 A x-ray structure of space-grown collagenase crystals.
Broutin-L'Hermite, I., Ries-Kautt, M., Ducruix, A.(2000) Acta Crystallogr D Biol Crystallogr 56: 376-378
- PubMed: 10713532
- DOI: https://doi.org/10.1107/s0907444999016789
- Primary Citation Related Structures:
2HLC - PubMed Abstract:
Collagenases, divided into metallocollagenases and serine collagenases, are the only proteases that cleave the triple helix of collagen under physiological conditions. In the present work, the serine protease collagenase purified from Hypoderma lineatum larvae is studied. From crystals grown in the International Microgravity Laboratory (IML2), a data set was collected at 1.7 A using synchrotron radiation. Although the resolution is not very different, the signal-to-noise ratio and the quality of the electron density are much improved. Alternate conformations were revealed for several residues, in particular Tyr99, suggesting a gate mechanism of recognition.
- Laboratoire d'Enzymologie et de Biochimie Structurales, Bâtiment 34, CNRS, 91198 Gif sur Yvette CEDEX, France. broutin@lebs.cnrs-gif.fr
Organizational Affiliation: