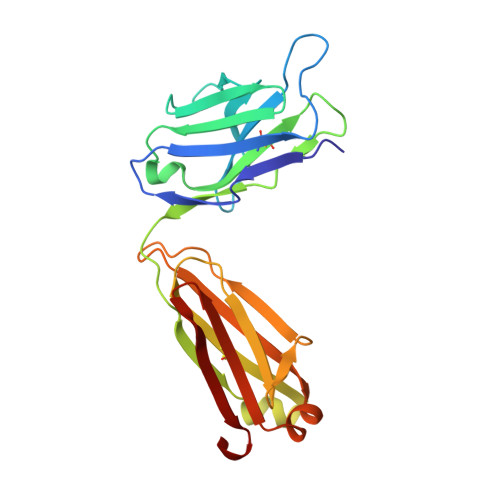
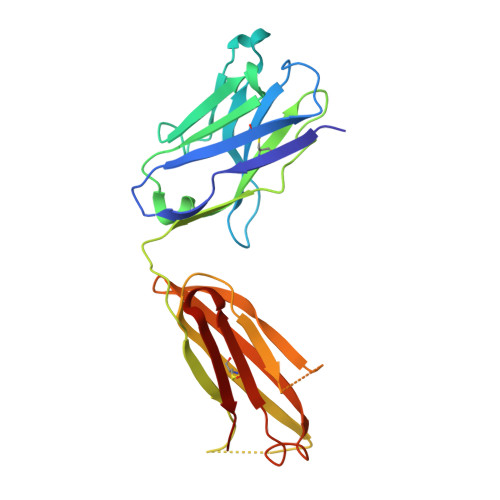
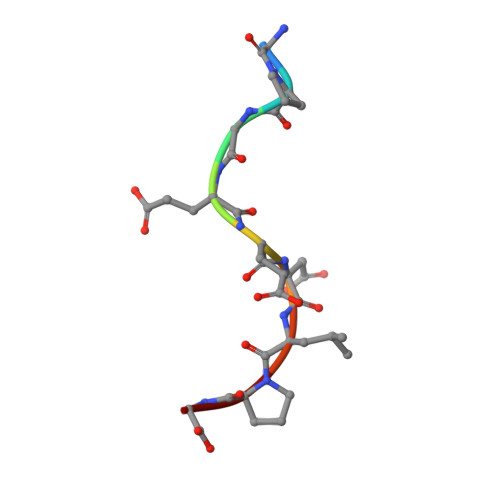
Stabilization of antibody structure upon association to a human carbonic anhydrase IX epitope studied by X-ray crystallography, microcalorimetry, and molecular dynamics simulations.
Kral, V., Mader, P., Collard, R., Fabry, M., Horejsi, M., Rezacova, P., Kozisek, M., Zavada, J., Sedlacek, J., Rulisek, L., Brynda, J.(2008) Proteins 71: 1275-1287
- PubMed: 18041760 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21821
- Primary Citation Related Structures:
2HKF, 2HKH - PubMed Abstract:
Specific antibodies interfere with the function of human tumor-associated carbonic anhydrase IX (CA IX), and show potential as tools for anticancer interventions. In this work, a correlation between structural elements and thermodynamic parameters of the association of antibody fragment Fab M75 to a peptide corresponding to its epitope in the proteoglycan-like domain of CA IX, is presented. Comparisons of the crystal structures of free Fab M75 and its complex with the epitope peptide reveal major readjustments of CDR-H1 and CDR-H3. In contrast, the overall conformations and positions of CDR-H2 and CDR-L2 remain unaltered, and their positively charged residues may thus present a fixed frame for epitope recognition. Adoption of the altered CDR-H3 conformation in the structure of the complex is accompanied by an apparent local stabilization. Analysis of domain mobility with translation-libration-screw (TLS) method shows that librations of the entire heavy chain variable domain (V(H)) decrease and reorient in the complex, which correlates well with participation of the heavy chain in ligand binding. Isothermal titration microcalorimetry (ITC) experiments revealed a highly unfavorable entropy term, which can be attributed mainly to the decrease in the degrees of freedom of the system, the loss of conformational freedom of peptide and partially to a local stabilization of CDR-H3. Moreover, it was observed that one proton is transferred from the environment to the protein-ligand complex upon binding. Molecular dynamics simulations followed by molecular mechanics/generalized Born surface area (MM-GBSA) calculations of the ligand (epitope peptide) binding energy yielded energy values that were in agreement with the ITC measurements and indicated that the charged residues play crucial role in the epitope binding. Theoretical arguments presented in this work indicate that two adjacent arginine residues (ArgH50 and ArgH52) are responsible for the observed proton transfer.
- Department of Recombinant Expression and Structural Biology, Institute of Molecular Genetics, Academy of Sciences of the Czech Republic, 166 37 Prague, Czech Republic.
Organizational Affiliation: