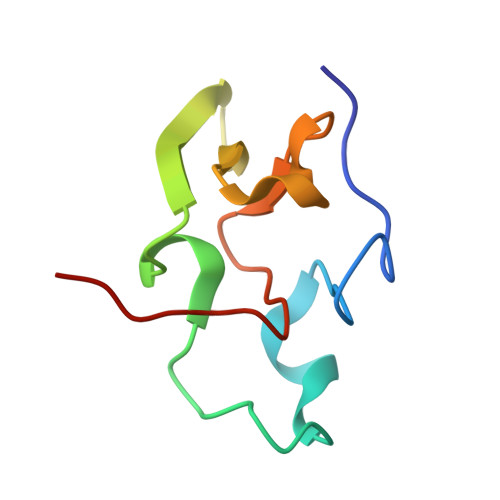
The molecular structure of the high potential iron-sulfur protein isolated from Ectothiorhodospira halophila determined at 2.5-A resolution.
Breiter, D.R., Meyer, T.E., Rayment, I., Holden, H.M.(1991) J Biological Chem 266: 18660-18667
- PubMed: 1917989 Search on PubMed
- DOI: https://doi.org/10.2210/pdb2hip/pdb
- Primary Citation Related Structures:
2HIP - PubMed Abstract:
The molecular structure of a high potential iron-sulfur protein (HiPIP) isolated from the purple photosynthetic bacterium, Ectothiorhodospira halophila strain BN9626, has been solved by x-ray diffraction analysis to a nominal resolution of 2.5 A and refined to a crystallographic R value of 18.4% including all measured x-ray data from 30.0- to 2.5-A resolution. Crystals used in the investigation contained two molecules/asymmetric unit and belonged to the space group P21 with unit cell dimensions of a = 60.00 A, b = 31.94 A, c = 40.27 A, and beta = 100.5 degrees. An interpretable electron density map, obtained by combining x-ray data from one isomorphous heavy atom derivative with non-crystallographic symmetry averaging and solvent flattening, clearly showed that this high potential iron-sulfur protein contains 71 amino acid residues, rather than 70 as originally reported. As in other bacterial ferredoxins, the [4Fe-4S] cluster adopts a cubane-like conformation and is ligated to the protein via four cysteinyl sulfur ligands. The overall secondary structure of the E. halophila HiPIP is characterized by a series of Type I and Type II turns allowing the polypeptide chain to wrap around the [4Fe-4S] prosthetic group. The hydrogen bonding pattern around the cluster is nearly identical to that originally observed in the 85-amino acid residue Chromatium vinosum HiPIP and consequently, the 240 mV difference in redox potentials between these two proteins cannot be simply attributed to hydrogen bonding patterns alone.
- Department of Chemistry, University of Wisconsin, Madison 53705.
Organizational Affiliation: