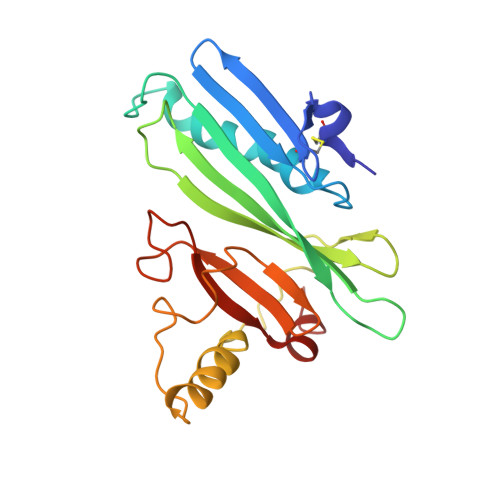
The Solution Structure of Antigen MPT64 from Mycobacterium tuberculosis Defines a New Family of Beta-Grasp Proteins.
Wang, Z., Potter, B.M., Gray, A.M., Sacksteder, K.A., Geisbrecht, B.V., Laity, J.H.(2007) J Mol Biology 366: 375-381
- PubMed: 17174329 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.11.039
- Primary Citation Related Structures:
2HHI - PubMed Abstract:
The MPT64 protein and its homologs form a highly conserved family of secreted proteins with unknown function that are found within the pathogenic Mycobacteria genus. The founding member of this family from Mycobacterium tuberculosis (MPT64 or protein Rv1980c) is expressed only when Mycobacteria cells are actively dividing. By virtue of this relatively unique expression profile, Rv1980c is currently under phase III clinical trials to evaluate its potential to replace tuberculin, or purified protein derivative, as the rapid diagnostic of choice for detection of active tuberculosis infection. We describe here the NMR solution structure of Rv1980c. This structure reveals a previously undescribed fold that is based upon a variation of a beta-grasp motif most commonly found in protein-protein interaction domains. Examination of this structure in conjunction with multiple sequence alignments of MPT64 homologs identifies a candidate ligand-binding site, which may help guide future studies of Rv1980c function. The work presented here also suggests structure-based approaches for increasing the antigenic potency of a Rv1980c-based diagnostic.
- Division of Cell Biology and Biophysics, School of Biological Sciences, University of Missouri-Kansas City, Kansas City, MO 64110-2499, USA.
Organizational Affiliation: