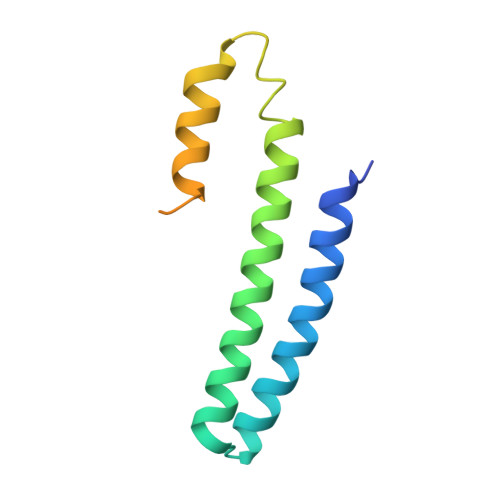
CsoR is a novel Mycobacterium tuberculosis copper-sensing transcriptional regulator.
Liu, T., Ramesh, A., Ma, Z., Ward, S.K., Zhang, L., George, G.N., Talaat, A.M., Sacchettini, J.C., Giedroc, D.P.(2007) Nat Chem Biol 3: 60-68
- PubMed: 17143269 Search on PubMed
- DOI: https://doi.org/10.1038/nchembio844
- Primary Citation Related Structures:
2HH7 - PubMed Abstract:
Copper is an essential element that becomes highly cytotoxic when concentrations exceed the capacity of cells to sequester the ion. Here, we identify a new copper-specific repressor (CsoR) of a copper-sensitive operon (cso) in Mycobacterium tuberculosis (Mtb) that is representative of a large, previously uncharacterized family of proteins (DUF156). Electronic and X-ray absorption spectroscopies reveal that CsoR binds a single-monomer mole equivalent of Cu(I) to form a trigonally coordinated (S(2)N) Cu(I) complex. The 2.6-A crystal structure of copper-loaded CsoR shows a homodimeric antiparallel four-helix bundle architecture that represents a novel DNA-binding fold. The Cu(I) is coordinated by Cys36, Cys65' and His61' in a subunit bridging site. Cu(I) binding negatively regulates the binding of CsoR to a DNA fragment encompassing the operator-promoter region of the Mtb cso operon; this results in derepression of the operon in Mtb and the heterologous host Mycobacterium smegmatis. Substitution of Cys36 or His61 with alanine abolishes Cu(I)- and CsoR-dependent regulation in vivo and in vitro. Potential roles of CsoR in Mtb pathogenesis are discussed.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, Texas 77843-2128, USA.
Organizational Affiliation: