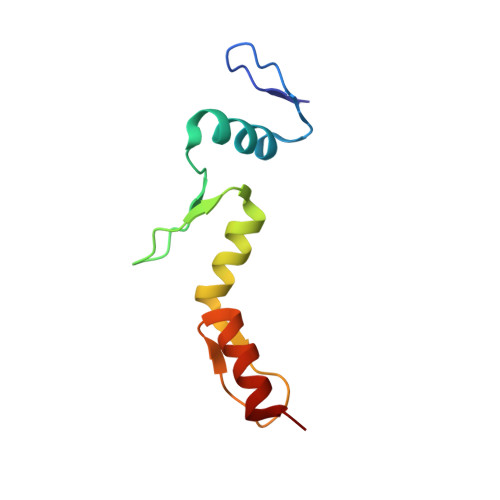
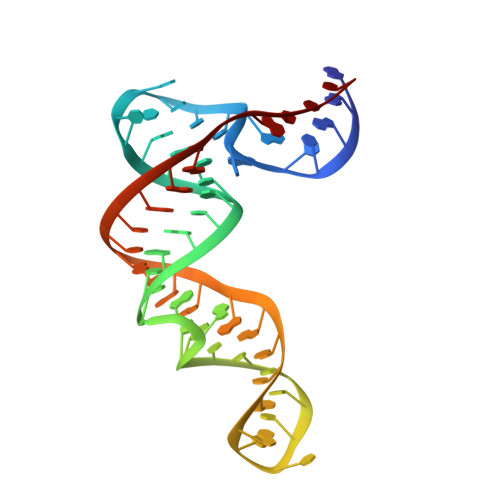
Induced Fit and 'Lock and Key' Recognition of 5 S RNA by Zinc Fingers of Transcription Factor IIIA
Lee, B.M., Xu, J., Clarkson, B.K., Martinez-Yamout, M.A., Dyson, J.H., Case, D.A., Gottesfeld, J.M., Wright, P.E.(2006) J Mol Biology 357: 275-291
- PubMed: 16405997 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.12.010
- Primary Citation Related Structures:
2HGH - PubMed Abstract:
Transcription factor IIIA (TFIIIA) is a Cys2His2 zinc finger protein that regulates expression of the 5 S ribosomal RNA gene by binding specifically to the internal control element. TFIIIA also functions in transport and storage of 5 S RNA by binding directly to the RNA transcript. To obtain insights into the mechanism by which TFIIIA recognizes 5 S RNA, we determined the solution structure of the middle three zinc fingers bound to the central core of 5 S RNA. Finger 4 utilizes "lock and key" recognition to bind in the widened major groove of the pre-structured RNA loop E motif. This interaction is mediated by direct hydrogen bonding interactions with bases. In contrast, recognition of loop A, a flexible junction of three helices, occurs by an induced fit mechanism that involves reorganization of the conserved CAUA motif and structuring of the finger 5-finger 6 interface to form a complementary RNA binding surface.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: