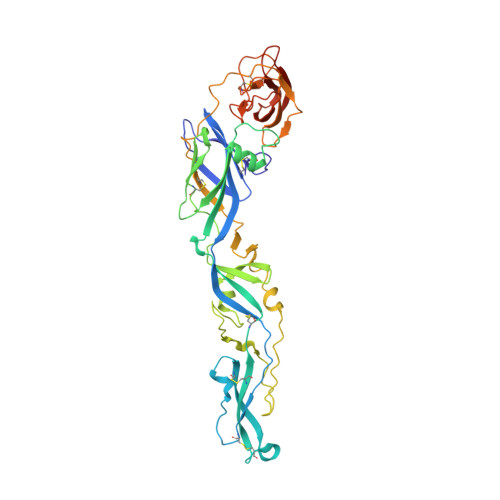
Crystal structure of the West Nile virus envelope glycoprotein.
Nybakken, G.E., Nelson, C.A., Chen, B.R., Diamond, M.S., Fremont, D.H.(2006) J Virol 80: 11467-11474
- PubMed: 16987985 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01125-06
- Primary Citation Related Structures:
2HG0 - PubMed Abstract:
The envelope glycoprotein (E) of West Nile virus (WNV) undergoes a conformational rearrangement triggered by low pH that results in a class II fusion event required for viral entry. Herein we present the 3.0-A crystal structure of the ectodomain of WNV E, which reveals insights into the flavivirus life cycle. We found that WNV E adopts a three-domain architecture that is shared by the E proteins from dengue and tick-borne encephalitis viruses and forms a rod-shaped configuration similar to that observed in immature flavivirus particles. Interestingly, the single N-linked glycosylation site on WNV E is displaced by a novel alpha-helix, which could potentially alter lectin-mediated attachment. The localization of histidines within the hinge regions of E implicates these residues in pH-induced conformational transitions. Most strikingly, the WNV E ectodomain crystallized as a monomer, in contrast to other flavivirus E proteins, which have crystallized as antiparallel dimers. WNV E assembles in a crystalline lattice of perpendicular molecules, with the fusion loop of one E protein buried in a hydrophobic pocket at the DI-DIII interface of another. Dimeric E proteins pack their fusion loops into analogous pockets at the dimer interface. We speculate that E proteins could pivot around the fusion loop-pocket junction, allowing virion conformational transitions while minimizing fusion loop exposure.
- Department of Pathology & Immunology, Washington University School of Medicine, Campus Box 8118, 660 South Euclid Avenue, St. Louis, MO 63110-1093, USA.
Organizational Affiliation: