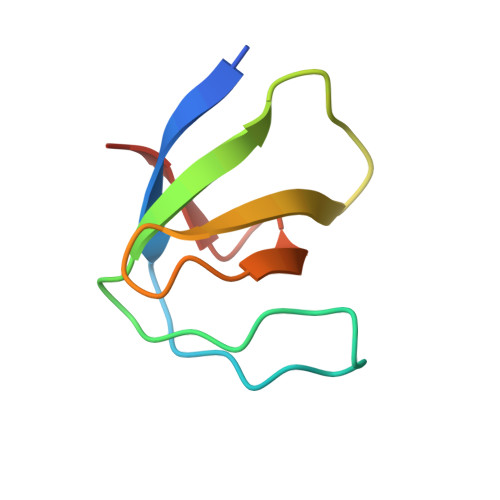
Crystallographic structure of the SH3 domain of the human c-Yes tyrosine kinase: Loop flexibility and amyloid aggregation.
Martin-Garcia, J.M., Luque, I., Mateo, P.L., Ruiz-Sanz, J., Camara-Artigas, A.(2007) FEBS Lett 581: 1701-1706
- PubMed: 17418139 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2007.03.059
- Primary Citation Related Structures:
2HDA - PubMed Abstract:
SH3 domains from the Src family of tyrosine kinases represent an interesting example of the delicate balance between promiscuity and specificity characteristic of proline-rich ligand recognition by SH3 domains. The development of inhibitors of therapeutic potential requires a good understanding of the molecular determinants of binding affinity and specificity and relies on the availability of high quality structural information. Here, we present the first high-resolution crystal structure of the SH3 domain of the c-Yes oncogen. Comparison with other SH3 domains from the Src family revealed significant deviations in the loop regions. In particular, the n-Src loop, highly flexible and partially disordered, is stabilized in an unusual conformation by the establishment of several intramolecular hydrogen bonds. Additionally, we present here the first report of amyloid aggregation by an SH3 domain from the Src family.
- Department of Physical Chemistry and Institute of Biotechnology, Faculty of Sciences, University of Granada, 18071 Granada, Spain.
Organizational Affiliation: